Chemistry and structural biology of androgen receptor
- PMID: 16159155
- PMCID: PMC2096617
- DOI: 10.1021/cr020456u
Chemistry and structural biology of androgen receptor
Figures





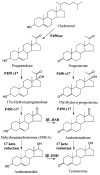

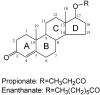
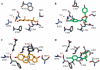
References
-
- Freedman LP. Molecular biology of steroid and nuclear hormone receptors. Birkhauser; Boston, MA: 1998.
-
- Mosselman S, Polman J, Dijkema R. FEBS Lett. 1996;392:49. - PubMed
-
- Fang H, Tong W, Branham WS, Moland CL, Dial SL, Hong H, Xie Q, Perkins R, Owens W, Sheehan DM. Chem. Res. Toxicol. 2003;16:1338. - PubMed
-
- Burger A, Abraham DJ. Burger's medicinal chemistry and drug discovery. 6th ed. Wiley; Hoboken, NJ: 2003.
-
- Foye WO, Williams DA, Lemke TL. Foye's principles of medicinal chemistry. 5th ed. Lippincott Williams & Wilkins; Philadelphia, PA: 2002.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

