Cell entry mechanism of enzymatic bacterial colicins: porin recruitment and the thermodynamics of receptor binding
- PMID: 16166265
- PMCID: PMC1236540
- DOI: 10.1073/pnas.0503567102
Cell entry mechanism of enzymatic bacterial colicins: porin recruitment and the thermodynamics of receptor binding
Abstract
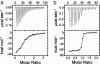
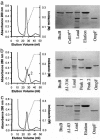
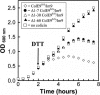
Binding of enzymatic E colicins to the vitamin B12 receptor, BtuB, is the first stage in a cascade of events that culminate in the translocation of the cytotoxic nuclease into the Escherichia coli cytoplasm and release of its tightly bound immunity protein. A dogma of colicin biology is that the toxin coiled-coil connecting its functional domains must unfold or unfurl to span the periplasm, with recent reports claiming this reaction is initiated by receptor binding. We report isothermal titration calorimetry data of BtuB binding the endonuclease toxin ColE9 and a disulfide form (ColE9S-S) where unfolding of the coiled-coil is prevented and, as a consequence, the toxin is biologically inactive. Contrary to expectation, the thermodynamics of receptor binding, characterized by large negative values for TDeltaS, are identical for the two colicins, arguing against any form of BtuB-induced unfolding. We go on to delineate key features of the "colicin translocon" that assembles at the cell surface after BtuB binding by using a complex of histidine-tagged Im9 bound to ColE9S-S. First, we show that the porin OmpF is recruited directly to the BtuB.colicin complex to form the translocon. Second, recruitment is through the natively unfolded region of the colicin translocation domain, with this domain likely having two contact points for OmpF. Finally, the immunity protein is not released during its assembly. Our study demonstrates that although colicin unfolding is undoubtedly a prerequisite for E. coli cell death, it must occur after assembly of the translocon.
Figures

References
-
- Falnes, P. O. & Sandvig, K. (2000) Curr. Opin. Cell Biol. 12, 407–413. - PubMed
-
- James, R., Lazdunski, C. J. & Pattus, F. (1992) Bacteriocins, Microcins, and Lantibiotics (Springer, Heidelberg, Germany).
-
- Brook, I. (1999) Crit. Rev. Microbiol. 25, 155–172. - PubMed
-
- Kirkup, B. C. & Riley, M. A. (2004) Nature 428, 412–414. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

