RNA silencing of single and multiple members in a gene family of rice
- PMID: 16172097
- PMCID: PMC1183382
- DOI: 10.1104/pp.105.063933
RNA silencing of single and multiple members in a gene family of rice
Abstract
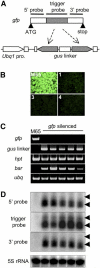
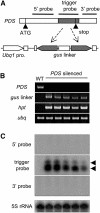
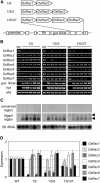
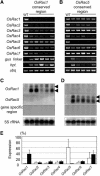
RNA silencing with inverted repeat (IR) constructs has been used to suppress gene expression in various organisms. However, the transitive RNA-silencing effect described in plants may preclude the use of RNA silencing for a gene family. Here, we show that, in rice (Oryza sativa), transitive RNA silencing (spreading of double-stranded RNA along the target mRNA) occurred with the green fluorescent protein transgene but not with the endogenous phytoene desaturase gene. We fused IR copies of unique 3' untranslated regions derived from the rice OsRac gene family to a strong promoter and stably introduced them into rice. Each of the seven members of the OsRac gene family was specifically suppressed by its respective IR construct. We also examined IR constructs in which multiple 3' untranslated regions were fused and showed that three members of the OsRac gene family were effectively suppressed by a single construct. Using highly conserved regions of the two members of the OsRac gene family, we also suppressed the expression of all members of the gene family with variable efficiencies. These results suggest that RNA silencing is a useful method for the functional analysis of gene families in rice and other plants.
Figures


References
-
- Azevedo C, Sadanandom A, Kitagawa K, Freialdenhoven A, Shirasu K, Schulze-Lefert P (2002) The RAR1 interactor SGT1, an essential component of R gene-triggered disease resistance. Science 295: 2073–2076 - PubMed
-
- Baulcombe D (2004) RNA silencing in plants. Nature 431: 356–363 - PubMed
-
- Beclin C, Boutet S, Waterhouse P, Vaucheret H (2002) A branched pathway for transgene-induced RNA silencing in plants. Curr Biol 12: 684–688 - PubMed
-
- Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409: 363–366 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

