Molecular signature of clinical severity in recovering patients with severe acute respiratory syndrome coronavirus (SARS-CoV)
- PMID: 16174304
- PMCID: PMC1262710
- DOI: 10.1186/1471-2164-6-132
Molecular signature of clinical severity in recovering patients with severe acute respiratory syndrome coronavirus (SARS-CoV)
Abstract
Background: Severe acute respiratory syndrome (SARS), a recent epidemic human disease, is caused by a novel coronavirus (SARS-CoV). First reported in Asia, SARS quickly spread worldwide through international travelling. As of July 2003, the World Health Organization reported a total of 8,437 people afflicted with SARS with a 9.6% mortality rate. Although immunopathological damages may account for the severity of respiratory distress, little is known about how the genome-wide gene expression of the host changes under the attack of SARS-CoV.
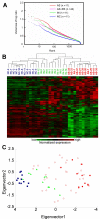
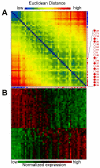
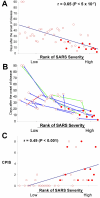
Results: Based on changes in gene expression of peripheral blood, we identified 52 signature genes that accurately discriminated acute SARS patients from non-SARS controls. While a general suppression of gene expression predominated in SARS-infected blood, several genes including those involved in innate immunity, such as defensins and eosinophil-derived neurotoxin, were upregulated. Instead of employing clustering methods, we ranked the severity of recovering SARS patients by generalized associate plots (GAP) according to the expression profiles of 52 signature genes. Through this method, we discovered a smooth transition pattern of severity from normal controls to acute SARS patients. The rank of SARS severity was significantly correlated with the recovery period (in days) and with the clinical pulmonary infection score.
Conclusion: The use of the GAP approach has proved useful in analyzing the complexity and continuity of biological systems. The severity rank derived from the global expression profile of significantly regulated genes in patients may be useful for further elucidating the pathophysiology of their disease.
Figures

Similar articles
-
A human in vitro model system for investigating genome-wide host responses to SARS coronavirus infection.BMC Infect Dis. 2004 Sep 9;4:34. doi: 10.1186/1471-2334-4-34. BMC Infect Dis. 2004. PMID: 15357874 Free PMC article.
-
[SARS-associated coronavirus gene fragments were detected from a suspected pediatric SARS patient].Zhonghua Er Ke Za Zhi. 2003 Sep;41(9):641-4. Zhonghua Er Ke Za Zhi. 2003. PMID: 14733796 Chinese.
-
Molecular diagnosis of severe acute respiratory syndrome.Methods Mol Biol. 2006;336:163-75. doi: 10.1385/1-59745-074-X:163. Methods Mol Biol. 2006. PMID: 16916262 Free PMC article.
-
Understanding the accessory viral proteins unique to the severe acute respiratory syndrome (SARS) coronavirus.Antiviral Res. 2006 Nov;72(2):78-88. doi: 10.1016/j.antiviral.2006.05.010. Epub 2006 Jun 6. Antiviral Res. 2006. PMID: 16820226 Free PMC article. Review.
-
The molecular biology of SARS coronavirus.Ann N Y Acad Sci. 2007 Apr;1102(1):26-38. doi: 10.1196/annals.1408.002. Ann N Y Acad Sci. 2007. PMID: 17470909 Free PMC article. Review.
Cited by
-
Immunopathogenesis of Different Emerging Viral Infections: Evasion, Fatal Mechanism, and Prevention.Front Immunol. 2021 Jul 15;12:690976. doi: 10.3389/fimmu.2021.690976. eCollection 2021. Front Immunol. 2021. PMID: 34335596 Free PMC article. Review.
-
Methods for simultaneously identifying coherent local clusters with smooth global patterns in gene expression profiles.BMC Bioinformatics. 2008 Mar 20;9:155. doi: 10.1186/1471-2105-9-155. BMC Bioinformatics. 2008. PMID: 18366693 Free PMC article.
-
Identification of genes associated with tumor development in CaSki cells in the cosmic space.Mol Biol Rep. 2012 Jun;39(6):6923-31. doi: 10.1007/s11033-012-1519-x. Mol Biol Rep. 2012. PMID: 22302396
-
Eosinopenia as Predictor of Poor Outcome in Hospitalized COVID-19 Adult Patients from Waves 1 and 2 of 2020 Pandemic.Microorganisms. 2022 Dec 7;10(12):2423. doi: 10.3390/microorganisms10122423. Microorganisms. 2022. PMID: 36557676 Free PMC article.
-
The successes and future prospects of the linear antisense RNA amplification methodology.Nat Protoc. 2018 May;13(5):811-818. doi: 10.1038/nprot.2018.011. Epub 2018 Mar 29. Nat Protoc. 2018. PMID: 29599441 Free PMC article.
References
-
- Rota PA, Oberste MS, Monroe SS, Nix WA, Campagnoli R, Icenogle JP, Penaranda S, Bankamp B, Maher K, Chen MH, Tong S, Tamin A, Lowe L, Frace M, DeRisi JL, Chen Q, Wang D, Erdman DD, Peret TC, Burns C, Ksiazek TG, Rollin PE, Sanchez A, Liffick S, Holloway B, Limor J, McCaustland K, Olsen-Rasmussen M, Fouchier R, Gunther S, Osterhaus AD, Drosten C, Pallansch MA, Anderson LJ, Bellini WJ. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science. 2003;300:1394–1399. doi: 10.1126/science.1085952. - DOI - PubMed
-
- Marra MA, Jones SJ, Astell CR, Holt RA, Brooks-Wilson A, Butterfield YS, Khattra J, Asano JK, Barber SA, Chan SY, Cloutier A, Coughlin SM, Freeman D, Girn N, Griffith OL, Leach SR, Mayo M, McDonald H, Montgomery SB, Pandoh PK, Petrescu AS, Robertson AG, Schein JE, Siddiqui A, Smailus DE, Stott JM, Yang GS, Plummer F, Andonov A, Artsob H, Bastien N, Bernard K, Booth TF, Bowness D, Czub M, Drebot M, Fernando L, Flick R, Garbutt M, Gray M, Grolla A, Jones S, Feldmann H, Meyers A, Kabani A, Li Y, Normand S, Stroher U, Tipples GA, Tyler S, Vogrig R, Ward D, Watson B, Brunham RC, Krajden M, Petric M, Skowronski DM, Upton C, Roper RL. The Genome sequence of the SARS-associated coronavirus. Science. 2003;300:1399–1404. doi: 10.1126/science.1085953. - DOI - PubMed
-
- Ruan YJ, Wei CL, Ee AL, Vega VB, Thoreau H, Su ST, Chia JM, Ng P, Chiu KP, Lim L, Zhang T, Peng CK, Lin EO, Lee NM, Yee SL, Ng LF, Chee RE, Stanton LW, Long PM, Liu ET. Comparative full-length genome sequence analysis of 14 SARS coronavirus isolates and common mutations associated with putative origins of infection. Lancet. 2003;361:1779–1785. doi: 10.1016/S0140-6736(03)13414-9. - DOI - PMC - PubMed
-
- Drosten C, Gunther S, Preiser W, van der Werf S, Brodt HR, Becker S, Rabenau H, Panning M, Kolesnikova L, Fouchier RA, Berger A, Burguiere AM, Cinatl J, Eickmann M, Escriou N, Grywna K, Kramme S, Manuguerra JC, Muller S, Rickerts V, Sturmer M, Vieth S, Klenk HD, Osterhaus AD, Schmitz H, Doerr HW. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med. 2003;348:1967–1976. doi: 10.1056/NEJMoa030747. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

