Dissection of the carbohydrate specificity of the broadly neutralizing anti-HIV-1 antibody 2G12
- PMID: 16174734
- PMCID: PMC1224641
- DOI: 10.1073/pnas.0505763102
Dissection of the carbohydrate specificity of the broadly neutralizing anti-HIV-1 antibody 2G12
Abstract
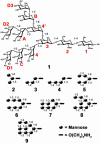
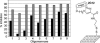
Human antibody 2G12 neutralizes a broad range of HIV-1 isolates. Hence, molecular characterization of its epitope, which corresponds to a conserved cluster of oligomannoses on the viral envelope glycoprotein gp120, is a high priority in HIV vaccine design. A prior crystal structure of 2G12 in complex with Man(9)GlcNAc(2) highlighted the central importance of the D1 arm in antibody binding. To characterize the specificity of 2G12 more precisely, we performed solution-phase ELISA, carbohydrate microarray analysis, and cocrystallized Fab 2G12 with four different oligomannose derivatives (Man(4), Man(5), Man(7), and Man(8)) that compete with gp120 for binding to 2G12. Our combined studies reveal that 2G12 is capable of binding both the D1 and D3 arms of the Man(9)GlcNAc(2) moiety, which would provide more flexibility to make the required multivalent interactions between the antibody and the gp120 oligomannose cluster than thought previously. These results have important consequences for the design of immunogens to elicit 2G12-like neutralizing antibodies as a component of an HIV vaccine.
Figures


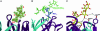
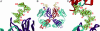
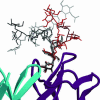
References
-
- Letvin, N. L. & Walker, B. D. (2003) Nat. Med. 9, 861-866. - PubMed
-
- McMichael, A. J. & Hanke, T. (2003) Nat. Med. 9, 874-880. - PubMed
-
- Burton, D. R., Desrosiers, R. C., Doms, R. W., Koff, W. C., Kwong, P. D., Moore, J. P., Nabel, G. J., Sodroski, J., Wilson, I. A. & Wyatt, R. T. (2004) Nat. Immunol. 5, 233-236. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

