Specific antibody can prevent fungal biofilm formation and this effect correlates with protective efficacy
- PMID: 16177306
- PMCID: PMC1230912
- DOI: 10.1128/IAI.73.10.6350-6362.2005
Specific antibody can prevent fungal biofilm formation and this effect correlates with protective efficacy
Abstract
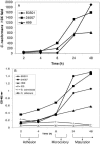
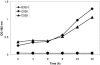
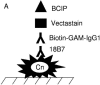
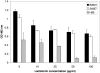
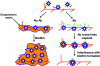
One of the most troublesome medical problems today is infection of prosthetic devices with organisms that form polysaccharide biofilms. This combined with increasing antimicrobial drug resistance is making many infectious diseases incurable. Cryptococcus neoformans is a human-pathogenic fungus that has a polysaccharide capsule and can form biofilms in prosthetic medical devices. We developed a system to study cryptococcal biofilm formation in vitro and studied the effect of antibody to the C. neoformans capsular polysaccharide on this process. C. neoformans biofilm formation was dependent on the presence of a polysaccharide capsule and correlated with the ability of capsular polysaccharide to bind the polystyrene solid support. Protective antibodies prevented biofilm formation whereas nonprotective antibodies were not effective. The mechanism of antibody action involved interference with capsular polysaccharide release from the fungal cell. In contrast, lactoferrin, an effector molecule of innate immune mechanisms, was unable to prevent fungal biofilm formation despite its efficacy against bacterial biofilms. Our results suggest a new role of adaptive humoral immunity whereby some antibodies can inhibit biofilm formation by encapsulated organisms. Vaccines that elicit antibody responses to capsular antigens and/or passive transfer of antibodies to microbial polysaccharides may be useful in preventing biofilm formation.
Figures









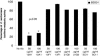
Similar articles
-
Specific antibody to Cryptococcus neoformans glucurunoxylomannan antagonizes antifungal drug action against cryptococcal biofilms in vitro.J Infect Dis. 2006 Jul 15;194(2):261-6. doi: 10.1086/504722. Epub 2006 Jun 14. J Infect Dis. 2006. PMID: 16779734
-
Antibody-mediated immobilization of Cryptococcus neoformans promotes biofilm formation.Appl Environ Microbiol. 2009 Apr;75(8):2528-33. doi: 10.1128/AEM.02846-08. Epub 2009 Feb 27. Appl Environ Microbiol. 2009. PMID: 19251903 Free PMC article.
-
Cryptococcus neoformans biofilm formation depends on surface support and carbon source and reduces fungal cell susceptibility to heat, cold, and UV light.Appl Environ Microbiol. 2007 Jul;73(14):4592-601. doi: 10.1128/AEM.02506-06. Epub 2007 May 18. Appl Environ Microbiol. 2007. PMID: 17513597 Free PMC article.
-
Cryptococcus neoformans: paradigm for the role of antibody immunity against fungi?Zentralbl Bakteriol. 1996 Aug;284(4):475-95. doi: 10.1016/s0934-8840(96)80001-6. Zentralbl Bakteriol. 1996. PMID: 8899968 Review.
-
Mechanism of action of antibody to capsular polysaccharide in Cryptococcus neoformans infection.Front Biosci. 1998 Feb 1;3:d136-51. doi: 10.2741/a270. Front Biosci. 1998. PMID: 9445465 Review.
Cited by
-
Immunoglobulins in defense, pathogenesis, and therapy of fungal diseases.Cell Host Microbe. 2012 May 17;11(5):447-56. doi: 10.1016/j.chom.2012.04.004. Cell Host Microbe. 2012. PMID: 22607798 Free PMC article. Review.
-
Susceptibility of Cryptococcus neoformans biofilms to antifungal agents in vitro.Antimicrob Agents Chemother. 2006 Mar;50(3):1021-33. doi: 10.1128/AAC.50.3.1021-1033.2006. Antimicrob Agents Chemother. 2006. PMID: 16495265 Free PMC article.
-
Anti-Immune Strategies of Pathogenic Fungi.Front Cell Infect Microbiol. 2016 Nov 15;6:142. doi: 10.3389/fcimb.2016.00142. eCollection 2016. Front Cell Infect Microbiol. 2016. PMID: 27896220 Free PMC article. Review.
-
Landmark clinical observations and immunopathogenesis pathways linked to HIV and Cryptococcus fatal central nervous system co-infection.Mycoses. 2020 Aug;63(8):840-853. doi: 10.1111/myc.13122. Epub 2020 Jun 19. Mycoses. 2020. PMID: 32472727 Free PMC article. Review.
-
Pathogenicity and virulence of Cryptococcus neoformans from an environmental perspective.Virulence. 2025 Dec;16(1):2547090. doi: 10.1080/21505594.2025.2547090. Epub 2025 Aug 14. Virulence. 2025. PMID: 40810603 Free PMC article. Review.
References
-
- Bullitt, R., and L. Makowski. 1995. Structural polymorphism of bacterial adhesion pili. Nature 373:164-167. - PubMed
-
- Casadevall, A., W. Cleare, M. Feldmesser, A. Glatman-Freedman, D. L. Goldman, T. R. Kozel, N. Lendvai, J. Mukherjee, L. A. Pirofski, J. Rivera, A. L. Rosas, M. D. Scharff, P. Valadon, K. Westin, and Z. Zhong. 1998. Characterization of a murine monoclonal antibody to Cryptococcus neoformans polysaccharide that is a candidate for human therapeutic studies. Antimicrob. Agents Chemother. 42:1437-1446. - PMC - PubMed
-
- Casadevall, A., J. Mukherjee, and M. D. Scharff. 1992. Monoclonal antibody based ELISAs for cryptococcal polysaccharide. J. Immunol. Methods 154:27-35. - PubMed
-
- Casadevall, A., and L. Pirofski. 2003. Antibody-mediated regulation of cellular immunity and the inflammatory response. Trends Immunol. 24:474-478. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

