Combining SELEX with quantitative assays to rapidly obtain accurate models of protein-DNA interactions
- PMID: 16186128
- PMCID: PMC1236725
- DOI: 10.1093/nar/gni139
Combining SELEX with quantitative assays to rapidly obtain accurate models of protein-DNA interactions
Abstract
Models for the specificity of DNA-binding transcription factors are often based on small amounts of qualitative data and therefore have limited accuracy. In this study we demonstrate a simple and efficient method of affinity chromatography-SELEX followed by a quantitative binding (QuMFRA) assay to rapidly collect the data necessary for more accurate models. Using the zinc finger protein EGR as an e.g. we show that many bindings sites can be obtained efficiently with affinity chromatography-SELEX, but those sequences alone provide a weight matrix model with limited accuracy. Using a QuMFRA assay to determine the quantitative relative affinity for only a subset of the sequences obtained by SELEX leads to a much more accurate model. Application of this method to variants of a transcription factor would allow us to generate a large collection of quantitative data for modeling protein-DNA interactions that could facilitate the determination of recognition codes for different transcription factor families.
Figures


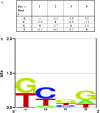

References
-
- Stormo G.D. DNA binding sites: representation and discovery. Bioinformatics. 2000;16:16–23. - PubMed
-
- Alexander M.K., Bourns B.D., Zakian V.A. One-hybrid systems for detecting protein–DNA interactions. Methods Mol. Biol. 2001;177:241–259. - PubMed
-
- Ren B., Robert F., Wyrick J.J., Aparicio O., Jennings E.G., Simon I., Zeitlinger J., Schreiber J., Hannett N., Kanin E., et al. Genome-wide location and function of DNA binding proteins. Science. 2000;290:2306–2309. - PubMed

