Critical role of the ubiquitin ligase activity of UHRF1, a nuclear RING finger protein, in tumor cell growth
- PMID: 16195352
- PMCID: PMC1289407
- DOI: 10.1091/mbc.e05-03-0194
Critical role of the ubiquitin ligase activity of UHRF1, a nuclear RING finger protein, in tumor cell growth
Abstract
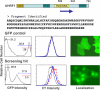
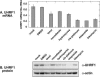
Early cellular events associated with tumorigenesis often include loss of cell cycle checkpoints or alteration in growth signaling pathways. Identification of novel genes involved in cellular proliferation may lead to new classes of cancer therapeutics. By screening a tetracycline-inducible cDNA library in A549 cells for genes that interfere with proliferation, we have identified a fragment of UHRF1 (ubiquitin-like protein containing PHD and RING domains 1), a nuclear RING finger protein, that acts as a dominant negative effector of cell growth. Reduction of UHRF1 levels using an UHRF1-specific shRNA decreased growth rates in several tumor cell lines. In addition, treatment of A549 cells with agents that activated different cell cycle checkpoints resulted in down-regulation of UHRF1. The primary sequence of UHRF1 contains a PHD and a RING motif, both of which are structural hallmarks of ubiquitin E3 ligases. We have confirmed using an in vitro autoubiquitination assay that UHRF1 displays RING-dependent E3 ligase activity. Overexpression of a GFP-fused UHRF1 RING mutant that lacks ligase activity sensitizes cells to treatment with various chemotherapeutics. Taken together, our results suggest a general requirement for UHRF1 in tumor cell proliferation and implicate the RING domain of UHRF1 as a functional determinant of growth regulation.
Figures


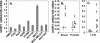




References
-
- Arima, Y., Hirota, T., Bronner, C., Mousli, M., Fujiwara, T., Niwa, S., Ishikawa, H., and Saya, H. (2004). Down-regulation of nuclear protein ICBP90 by p53/p21Cip1/WAF1-dependent DNA-damage checkpoint signals contributes to cell cycle arrest at G1/S transition. Genes Cells 9, 131–142. - PubMed
-
- Coscoy, L., and Ganem, D. (2003). PHD domains and E3 ubiquitin ligases: viruses make the connection. Trends Cell Biol. 13, 7–12. - PubMed
-
- Elbashir, S. M., Harborth, J., Lendeckel, W., Yalcin, A., Weber, K., and Tuschl, T. (2001). Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

