YrxA is the transcriptional regulator that represses de novo NAD biosynthesis in Bacillus subtilis
- PMID: 16199587
- PMCID: PMC1251630
- DOI: 10.1128/JB.187.20.7155-7160.2005
YrxA is the transcriptional regulator that represses de novo NAD biosynthesis in Bacillus subtilis
Abstract
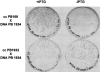
The first genetic, in vivo, and in vitro evidences that YrxA is the regulator of NAD de novo biosynthesis in Bacillus subtilis are hereby reported. The protein is essential to the transcription repression of the divergent operons nadBCA and nifS-yrxA in the presence of nicotinic acid and binds to their shared operator-promoter region.
Figures


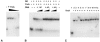
Similar articles
-
Transcriptional regulation of NAD metabolism in bacteria: genomic reconstruction of NiaR (YrxA) regulon.Nucleic Acids Res. 2008 Apr;36(6):2032-46. doi: 10.1093/nar/gkn046. Epub 2008 Feb 14. Nucleic Acids Res. 2008. PMID: 18276644 Free PMC article.
-
Cloning, nucleotide sequence, and regulation of the Bacillus subtilis nadB gene and a nifS-like gene, both of which are essential for NAD biosynthesis.J Bacteriol. 1993 Mar;175(5):1423-32. doi: 10.1128/jb.175.5.1423-1432.1993. J Bacteriol. 1993. PMID: 8444804 Free PMC article.
-
Bacillus subtilis ilvB operon: an intersection of global regulons.Mol Microbiol. 2005 Jun;56(6):1549-59. doi: 10.1111/j.1365-2958.2005.04634.x. Mol Microbiol. 2005. PMID: 15916605
-
Organization and regulation of genes for de novo purine nucleotide synthesis in Bacillus subtilis.Res Microbiol. 1991 Sep-Oct;142(7-8):765-9. doi: 10.1016/0923-2508(91)90053-d. Res Microbiol. 1991. PMID: 1784814 Review. No abstract available.
-
Systematic function analysis of Bacillus subtilis genes.Res Microbiol. 2000 Mar;151(2):129-34. doi: 10.1016/s0923-2508(00)00118-2. Res Microbiol. 2000. PMID: 10865958 Review.
Cited by
-
Transcriptional regulation of NAD metabolism in bacteria: NrtR family of Nudix-related regulators.Nucleic Acids Res. 2008 Apr;36(6):2047-59. doi: 10.1093/nar/gkn047. Epub 2008 Feb 14. Nucleic Acids Res. 2008. PMID: 18276643 Free PMC article.
-
Transcriptional regulation of NAD metabolism in bacteria: genomic reconstruction of NiaR (YrxA) regulon.Nucleic Acids Res. 2008 Apr;36(6):2032-46. doi: 10.1093/nar/gkn046. Epub 2008 Feb 14. Nucleic Acids Res. 2008. PMID: 18276644 Free PMC article.
-
Biogenesis and Homeostasis of Nicotinamide Adenine Dinucleotide Cofactor.EcoSal Plus. 2009 Aug;3(2):10.1128/ecosalplus.3.6.3.10. doi: 10.1128/ecosalplus.3.6.3.10. EcoSal Plus. 2009. PMID: 26443758 Free PMC article.
-
Comparative genomic reconstruction of transcriptional regulatory networks in bacteria.Chem Rev. 2007 Aug;107(8):3467-97. doi: 10.1021/cr068309+. Epub 2007 Jul 18. Chem Rev. 2007. PMID: 17636889 Free PMC article. Review. No abstract available.
-
Regulation of the expression of genes involved in NAD de novo biosynthesis in Corynebacterium glutamicum.Appl Environ Microbiol. 2010 Aug;76(16):5488-95. doi: 10.1128/AEM.00906-10. Epub 2010 Jul 2. Appl Environ Microbiol. 2010. PMID: 20601509 Free PMC article.
References
-
- Anantharaman, V., E. V. Koonin, and L. Aravind. 2001. Regulatory potential, phyletic distribution and evolution of ancient, intracellular, small-molecule-binding domains. J. Mol. Biol. 307:1271-1292. - PubMed
-
- Begley, T. P., C. Kinsland, R. A. Mehl, A. Osterman, and P. Dorrestein. 2001. The biosynthesis of nicotinamide adenine dinucleotides in bacteria. Vitam. Horm. 61:103-119. - PubMed
-
- Ceciliani, F., T. Caramori, S. Ronchi, G. Tedeschi, M. Mortarino, and A. Galizzi. 2000. Cloning, overexpression, and purification of Escherichia coli quinolinate synthetase. Protein Expr. Purif. 18:64-70. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

