Analysis of genes involved in arsenic resistance in Corynebacterium glutamicum ATCC 13032
- PMID: 16204540
- PMCID: PMC1266000
- DOI: 10.1128/AEM.71.10.6206-6215.2005
Analysis of genes involved in arsenic resistance in Corynebacterium glutamicum ATCC 13032
Abstract
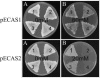
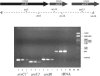
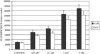
Corynebacterium glutamicum is able to grow in media containing up to 12 mM arsenite and 500 mM arsenate and is one of the most arsenic-resistant microorganisms described to date. Two operons (ars1 and ars2) involved in arsenate and arsenite resistance have been identified in the complete genome sequence of Corynebacterium glutamicum. The operons ars1 and ars2 are located some distance from each other in the bacterial chromosome, but they are both composed of genes encoding a regulatory protein (arsR), an arsenite permease (arsB), and an arsenate reductase (arsC); operon ars1 contains an additional arsenate reductase gene (arsC1') located immediately downstream from arsC1. Additional arsenite permease and arsenate reductase genes (arsB3 and arsC4) scattered on the chromosome were also identified. The involvement of ars operons in arsenic resistance in C. glutamicum was confirmed by gene disruption experiments of the three arsenite permease genes present in its genome. Wild-type and arsB3 insertional mutant C. glutamicum strains were able to grow with up to 12 mM arsenite, whereas arsB1 and arsB2 C. glutamicum insertional mutants were resistant to 4 mM and 9 mM arsenite, respectively. The double arsB1-arsB2 insertional mutant was resistant to only 0.4 mM arsenite and 10 mM arsenate. Gene amplification assays of operons ars1 and ars2 in C. glutamicum revealed that the recombinant strains containing the ars1 operon were resistant to up to 60 mM arsenite, this being one of the highest levels of bacterial resistance to arsenite so far described, whereas recombinant strains containing operon ars2 were resistant to only 20 mM arsenite. Northern blot and reverse transcription-PCR analysis confirmed the presence of transcripts for all the ars genes, the expression of arsB3 and arsC4 being constitutive, and the expression of arsR1, arsB1, arsC1, arsC1', arsR2, arsB2, and arsC2 being inducible by arsenite.
Figures

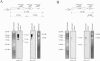
References
-
- Abdrashitova, S. A., G. G. Abdullina, and A. N. Ilialetdinov. 1986. Role of arsenites in lipid peroxidation in Pseudomonas putida cells oxidizing arsenite. Mikrobiologiya 55:212-216.
-
- Adham, S. A., S. Rodriguez, A. Ramos, R. I. Santamaria, and J. A. Gil. 2003. Improved vectors for transcriptional/translational signal screening in corynebacteria using the melC operon from Streptomyces glaucescens as reporter. Arch. Microbiol. 180:53-59. - PubMed
-
- Bhattacharjee, H., J. Li, M. Y. Ksenzenko, and B. P. Rosen. 1995. Role of cysteinyl residues in metalloactivation of the oxyanion-translocating ArsA ATPase. J. Biol. Chem. 270:11245-11250. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

