The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes
- PMID: 16214803
- PMCID: PMC1251668
- DOI: 10.1093/nar/gki866
The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes
Abstract
The release of the 1000th complete microbial genome will occur in the next two to three years. In anticipation of this milestone, the Fellowship for Interpretation of Genomes (FIG) launched the Project to Annotate 1000 Genomes. The project is built around the principle that the key to improved accuracy in high-throughput annotation technology is to have experts annotate single subsystems over the complete collection of genomes, rather than having an annotation expert attempt to annotate all of the genes in a single genome. Using the subsystems approach, all of the genes implementing the subsystem are analyzed by an expert in that subsystem. An annotation environment was created where populated subsystems are curated and projected to new genomes. A portable notion of a populated subsystem was defined, and tools developed for exchanging and curating these objects. Tools were also developed to resolve conflicts between populated subsystems. The SEED is the first annotation environment that supports this model of annotation. Here, we describe the subsystem approach, and offer the first release of our growing library of populated subsystems. The initial release of data includes 180 177 distinct proteins with 2133 distinct functional roles. This data comes from 173 subsystems and 383 different organisms.
Figures

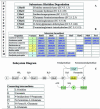
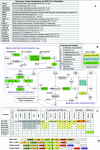

References
-
- Fleischmann R.D., Adams M.D., White O., Clayton R.A., Kirkness E.F., Kerlavage A.R., Bult C.J., Tomb J.F., Dougherty B.A., Merrick J.M., et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science. 1995;269:496–512. - PubMed
-
- Haft D.H., Selengut J.D., Brinkac L.M., Zafar N., White O. Genome Properties: a system for the investigation of prokaryotic genetic content for microbiology, genome annotation and comparative genomics. Bioinformatics. 2005;21:293–306. - PubMed
-
- Osterman A., Overbeek R. Missing genes in metabolic pathways: a comparative genomics approach. Curr. Opin. Chem. Biol. 2003;7:238–251. - PubMed
-
- Overbeek R., Larsen N., Smith W., Maltsev N., Selkov E. Representation of function: the next step. Gene. 1997;191:GC1–GC9. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

