RGS4 and RGS5 are in vivo substrates of the N-end rule pathway
- PMID: 16217033
- PMCID: PMC1257735
- DOI: 10.1073/pnas.0507533102
RGS4 and RGS5 are in vivo substrates of the N-end rule pathway
Abstract
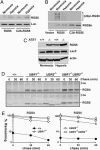
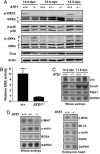
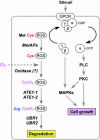
The ATE1-encoded Arg-transferase mediates conjugation of Arg to N-terminal Asp, Glu, and Cys of certain eukaryotic proteins, yielding N-terminal Arg that can act as a degradation signal for the ubiquitin-dependent N-end rule pathway. We have previously shown that mouse ATE1-/- embryos die with defects in heart development and angiogenesis. Here, we report that the ATE1 Arg-transferase mediates the in vivo degradation of RGS4 and RGS5, which are negative regulators of specific G proteins whose functions include cardiac growth and angiogenesis. The proteolysis of these regulators of G protein signaling (RGS) proteins was perturbed either by hypoxia or in cells lacking ubiquitin ligases UBR1 and/or UBR2. Mutant RGS proteins in which the conserved Cys-2 residue could not become N-terminal were long-lived in vivo. We propose a model in which the sequential modifications of RGS4, RGS5, and RGS16 (N-terminal exposure of their Cys-2, its oxidation, and subsequent arginylation) act as a licensing mechanism in response to extracellular and intracellular signals before the targeting for proteolysis by UBR1 and UBR2. We also show that ATE1-/- embryos are impaired in the activation of extracellular signal-regulated kinase mitogen-activated protein kinases and in the expression of G protein-induced downstream effectors such as Jun, cyclin D1, and beta-myosin heavy chain. These results establish RGS4 and RGS5 as in vivo substrates of the mammalian N-end rule pathway and also suggest that the O2-ATE1-UBR1/UBR2 proteolytic circuit plays a role in RGS-regulated G protein signaling in the cardiovascular system.
Figures


References
-
- Bachmair, A., Finley, D. & Varshavsky, A. (1986) Science 234, 179–186. - PubMed
-
- Kwon, Y. T., Kashina, A. S., Davydov, I. V., Hu, R. G., An, J. Y., Seo, J. W., Du, F. & Varshavsky, A. (2002) Science 297, 96–99. - PubMed
-
- Neves, S. R., Ram, P. T. & Iyengar, R. (2002) Science 296, 1636–1639. - PubMed
-
- Davydov, I. V. & Varshavsky, A. (2000) J. Biol. Chem. 275, 22931–22941. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

