Directional positive selection on an allele of arbitrary dominance
- PMID: 16219788
- PMCID: PMC1456198
- DOI: 10.1534/genetics.105.044065
Directional positive selection on an allele of arbitrary dominance
Abstract
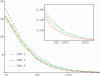
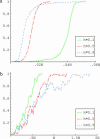
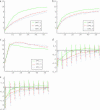
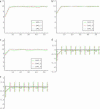
Most models of positive directional selection assume codominance of the beneficial allele. We examine the importance of this assumption by implementing a coalescent model of positive directional selection with arbitrary dominance. We find that, for a given mean fixation time, a beneficial allele has a much weaker effect on diversity at linked neutral sites when the allele is recessive.
Figures
References
-
- Barton, N. H., 1998. The effect of hitch-hiking on neutral genealogies. Genet. Res. 72: 123–133.
-
- Coop, G., and R. C. Griffiths, 2004. Ancestral inference on gene trees under selection. Theor. Popul. Biol. 66: 219–232. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

