Nonrandom packaging of host RNAs in moloney murine leukemia virus
- PMID: 16227273
- PMCID: PMC1262605
- DOI: 10.1128/JVI.79.21.13528-13537.2005
Nonrandom packaging of host RNAs in moloney murine leukemia virus
Abstract
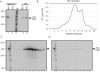
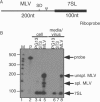
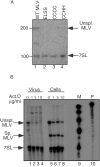
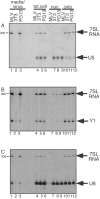
Moloney murine leukemia virus (MLV) particles contain both viral genomic RNA and an assortment of host cell RNAs. Packaging of virus-encoded RNA is selective, with virions virtually devoid of spliced env mRNA and highly enriched for unspliced genome. Except for primer tRNA, it is unclear whether packaged host RNAs are randomly sampled from the cell or specifically encapsidated. To address possible biases in host RNA sampling, the relative abundances of several host RNAs in MLV particles and in producer cells were compared. Using 7SL RNA as a standard, some cellular RNAs, such as those of the Ro RNP, were found to be enriched in MLV particles in that their ratios relative to 7SL differed little, if at all, from their ratios in cells. Some RNAs were underrepresented, with ratios relative to 7SL several orders of magnitude lower in virions than in cells, while others displayed intermediate values. At least some enriched RNAs were encapsidated by genome-defective nucleocapsid mutants. Virion RNAs were not a random sample of the cytosol as a whole, since some cytoplasmic RNAs like tRNA(Met) were vastly underrepresented, while U6 spliceosomal RNA, which functions in the nucleus, was enriched. Real-time PCR demonstrated that env mRNA, although several orders of magnitude less abundant than unspliced viral RNA, was slightly enriched relative to actin mRNA in virions. These data demonstrate that certain host RNAs are nearly as enriched in virions as genomic RNA and suggest that Psi- mRNAs and some other host RNAs may be specifically excluded from assembly sites.
Figures


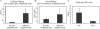
References
-
- Basyuk, E., T. Galli, M. Mougel, J.-M. Blanchard, M. Sitbon, and E. Bertrand. 2003. Retroviral genomic RNAs are transported to the plasma membrane by endosomal vesicles. Dev. Cell 5:161-174. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

