Nuclear dynamics of PCNA in DNA replication and repair
- PMID: 16227586
- PMCID: PMC1265825
- DOI: 10.1128/MCB.25.21.9350-9359.2005
Nuclear dynamics of PCNA in DNA replication and repair
Abstract
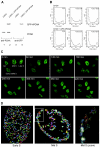
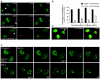
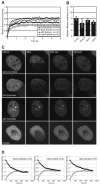
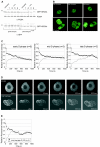
The DNA polymerase processivity factor proliferating cell nuclear antigen (PCNA) is central to both DNA replication and repair. The ring-shaped homotrimeric PCNA encircles and slides along double-stranded DNA, acting as a "sliding clamp" that localizes proteins to DNA. We determined the behavior of green fluorescent protein-tagged human PCNA (GFP-hPCNA) in living cells to analyze its different engagements in DNA replication and repair. Photobleaching and tracking of replication foci revealed a dynamic equilibrium between two kinetic pools of PCNA, i.e., bound to replication foci and as a free mobile fraction. To simultaneously monitor PCNA action in DNA replication and repair, we locally inflicted UV-induced DNA damage. A surprisingly longer residence time of PCNA at damaged areas than at replication foci was observed. Using DNA repair mutants, we showed that the initial recruitment of PCNA to damaged sites was dependent on nucleotide excision repair. Local accumulation of PCNA at damaged regions was observed during all cell cycle stages but temporarily disappeared during early S phase. The reappearance of PCNA accumulation in discrete foci at later stages of S phase likely reflects engagements of PCNA in distinct genome maintenance processes dealing with stalled replication forks, such as translesion synthesis (TLS). Using a ubiquitination mutant of GFP-hPCNA that is unable to participate in TLS, we noticed a significantly shorter residence time in damaged areas. Our results show that changes in the position of PCNA result from de novo assembly of freely mobile replication factors in the nucleoplasmic pool and indicate different binding affinities for PCNA in DNA replication and repair.
Figures

Similar articles
-
High mobility of flap endonuclease 1 and DNA polymerase eta associated with replication foci in mammalian S-phase nucleus.Mol Biol Cell. 2005 May;16(5):2518-28. doi: 10.1091/mbc.e04-12-1066. Epub 2005 Mar 9. Mol Biol Cell. 2005. PMID: 15758026 Free PMC article.
-
Temporally distinct translesion synthesis pathways for ultraviolet light-induced photoproducts in the mammalian genome.DNA Repair (Amst). 2012 Jun 1;11(6):550-8. doi: 10.1016/j.dnarep.2012.03.007. Epub 2012 Apr 20. DNA Repair (Amst). 2012. PMID: 22521143
-
Lys-110 is essential for targeting PCNA to replication and repair foci, and the K110A mutant activates apoptosis.Biol Cell. 2008 Dec;100(12):675-86. doi: 10.1042/BC20070158. Biol Cell. 2008. PMID: 18498247 Free PMC article.
-
PCNA modifications for regulation of post-replication repair pathways.Mol Cells. 2008 Jul 31;26(1):5-11. Epub 2008 Jun 4. Mol Cells. 2008. PMID: 18525240 Free PMC article. Review.
-
PCNA cycling dynamics during DNA replication and repair in mammals.Trends Genet. 2024 Jun;40(6):526-539. doi: 10.1016/j.tig.2024.02.006. Epub 2024 Mar 13. Trends Genet. 2024. PMID: 38485608 Review.
Cited by
-
Role of reduced manganese superoxide dismutase in ischemia-reperfusion injury: a possible trigger for autophagy and mitochondrial biogenesis?Am J Physiol Renal Physiol. 2013 Feb 1;304(3):F257-67. doi: 10.1152/ajprenal.00435.2012. Epub 2012 Nov 28. Am J Physiol Renal Physiol. 2013. PMID: 23195678 Free PMC article.
-
Cyclin Kinase-independent role of p21CDKN1A in the promotion of nascent DNA elongation in unstressed cells.Elife. 2016 Oct 14;5:e18020. doi: 10.7554/eLife.18020. Elife. 2016. PMID: 27740454 Free PMC article.
-
Glucans from the edible mushroom Pleurotus pulmonarius inhibit colitis-associated colon carcinogenesis in mice.J Gastroenterol. 2012 May;47(5):504-18. doi: 10.1007/s00535-011-0514-7. Epub 2011 Dec 21. J Gastroenterol. 2012. PMID: 22187166
-
Transcriptome reprogramming during severe dehydration contributes to physiological and metabolic changes in the resurrection plant Haberlea rhodopensis.BMC Plant Biol. 2018 Dec 13;18(1):351. doi: 10.1186/s12870-018-1566-0. BMC Plant Biol. 2018. PMID: 30541446 Free PMC article.
-
Sequential recruitment of the repair factors during NER: the role of XPG in initiating the resynthesis step.EMBO J. 2008 Jan 9;27(1):155-67. doi: 10.1038/sj.emboj.7601948. Epub 2007 Dec 13. EMBO J. 2008. PMID: 18079701 Free PMC article.
References
-
- Aboussekhra, A., and R. D. Wood. 1995. Detection of nucleotide excision repair incisions in human fibroblasts by immunostaining for PCNA. Exp. Cell Res. 221:326-332. - PubMed
-
- Chubb, J. R., S. Boyle, P. Perry, and W. A. Bickmore. 2002. Chromatin motion is constrained by association with nuclear compartments in human cells. Curr. Biol. 12:439-445. - PubMed
-
- Dunand-Sauthier, I., M. Hohl, F. Thorel, P. Jaquier-Gubler, S. G. Clarkson, and O. D. Scharer. 2005. The spacer region of XPG mediates recruitment to nucleotide excision repair complexes and determines substrate specificity. J. Biol. Chem. 280:7030-7037. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
