Generation, annotation, analysis and database integration of 16,500 white spruce EST clusters
- PMID: 16236172
- PMCID: PMC1277824
- DOI: 10.1186/1471-2164-6-144
Generation, annotation, analysis and database integration of 16,500 white spruce EST clusters
Abstract
Background: The sequencing and analysis of ESTs is for now the only practical approach for large-scale gene discovery and annotation in conifers because their very large genomes are unlikely to be sequenced in the near future. Our objective was to produce extensive collections of ESTs and cDNA clones to support manufacture of cDNA microarrays and gene discovery in white spruce (Picea glauca [Moench] Voss).
Results: We produced 16 cDNA libraries from different tissues and a variety of treatments, and partially sequenced 50,000 cDNA clones. High quality 3' and 5' reads were assembled into 16,578 consensus sequences, 45% of which represented full length inserts. Consensus sequences derived from 5' and 3' reads of the same cDNA clone were linked to define 14,471 transcripts. A large proportion (84%) of the spruce sequences matched a pine sequence, but only 68% of the spruce transcripts had homologs in Arabidopsis or rice. Nearly all the sequences that matched the Populus trichocarpa genome (the only sequenced tree genome) also matched rice or Arabidopsis genomes. We used several sequence similarity search approaches for assignment of putative functions, including blast searches against general and specialized databases (transcription factors, cell wall related proteins), Gene Ontology term assignation and Hidden Markov Model searches against PFAM protein families and domains. In total, 70% of the spruce transcripts displayed matches to proteins of known or unknown function in the Uniref100 database (blastx e-value < 1e-10). We identified multigenic families that appeared larger in spruce than in the Arabidopsis or rice genomes. Detailed analysis of translationally controlled tumour proteins and S-adenosylmethionine synthetase families confirmed a twofold size difference. Sequences and annotations were organized in a dedicated database, SpruceDB. Several search tools were developed to mine the data either based on their occurrence in the cDNA libraries or on functional annotations.
Conclusion: This report illustrates specific approaches for large-scale gene discovery and annotation in an organism that is very distantly related to any of the fully sequenced genomes. The ArboreaSet sequences and cDNA clones represent a valuable resource for investigations ranging from plant comparative genomics to applied conifer genetics.
Figures


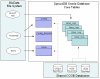

Similar articles
-
A conifer genomics resource of 200,000 spruce (Picea spp.) ESTs and 6,464 high-quality, sequence-finished full-length cDNAs for Sitka spruce (Picea sitchensis).BMC Genomics. 2008 Oct 14;9:484. doi: 10.1186/1471-2164-9-484. BMC Genomics. 2008. PMID: 18854048 Free PMC article.
-
Generation, functional annotation and comparative analysis of black spruce (Picea mariana) ESTs: an important conifer genomic resource.BMC Genomics. 2013 Oct 11;14:702. doi: 10.1186/1471-2164-14-702. BMC Genomics. 2013. PMID: 24119028 Free PMC article.
-
Identification of conserved core xylem gene sets: conifer cDNA microarray development, transcript profiling and computational analyses.New Phytol. 2008;180(4):766-86. doi: 10.1111/j.1469-8137.2008.02615.x. New Phytol. 2008. PMID: 18811621
-
Identification and analysis of gene families from the duplicated genome of soybean using EST sequences.BMC Genomics. 2006 Aug 9;7:204. doi: 10.1186/1471-2164-7-204. BMC Genomics. 2006. PMID: 16899135 Free PMC article. Review.
-
Large-scale EST sequencing in rice.Plant Mol Biol. 1997 Sep;35(1-2):135-44. Plant Mol Biol. 1997. PMID: 9291967 Review.
Cited by
-
A spruce gene map infers ancient plant genome reshuffling and subsequent slow evolution in the gymnosperm lineage leading to extant conifers.BMC Biol. 2012 Oct 26;10:84. doi: 10.1186/1741-7007-10-84. BMC Biol. 2012. PMID: 23102090 Free PMC article.
-
Comparative genome mapping among Picea glauca, P. mariana x P. rubens and P. abies, and correspondence with other Pinaceae.Theor Appl Genet. 2006 Nov;113(8):1371-93. doi: 10.1007/s00122-006-0354-7. Epub 2006 Oct 24. Theor Appl Genet. 2006. PMID: 17061103
-
Modern Approaches for Transcriptome Analyses in Plants.Adv Exp Med Biol. 2021;1346:11-50. doi: 10.1007/978-3-030-80352-0_2. Adv Exp Med Biol. 2021. PMID: 35113394
-
Identification and functional characterization of monofunctional ent-copalyl diphosphate and ent-kaurene synthases in white spruce reveal different patterns for diterpene synthase evolution for primary and secondary metabolism in gymnosperms.Plant Physiol. 2010 Mar;152(3):1197-208. doi: 10.1104/pp.109.151456. Epub 2009 Dec 31. Plant Physiol. 2010. PMID: 20044448 Free PMC article.
-
Comparative analysis of the small RNA transcriptomes of Pinus contorta and Oryza sativa.Genome Res. 2008 Apr;18(4):571-84. doi: 10.1101/gr.6897308. Epub 2008 Mar 6. Genome Res. 2008. PMID: 18323537 Free PMC article.
References
-
- Ahuja MR. Recent advances in molecular genetics of forest trees. Euphytica. 2001;121:173–195. doi: 10.1023/A:1012226319449. - DOI
-
- Dhillon SS. DNA in tree species. In: Bonga JM, Durzan DJ, editor. Cell and Tissue Culture in Forestry. Vol. 1. Martinus Nijhoff Publishers, Dordrecht; 1987. pp. 298–313.
-
- Wakamiya I, Newton RJ, Price JS. Genome size and environmental factors in the genus Pinus. Am J Bot. 1993;80:1235–1241.
-
- Rake AW, Miksche JP, Hall RB, Hanson KM. DNA reassociation kinetics for four conifers. Can J Genet Cytol. 1980;22:69–79.
-
- Ohri D, Khoshoo TN. Genome size in gymnosperms. Plant Syst Evol. 1986;153:119–132. doi: 10.1007/BF00989421. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

