Multiple and interconnected pathways for L-lysine catabolism in Pseudomonas putida KT2440
- PMID: 16237033
- PMCID: PMC1272968
- DOI: 10.1128/JB.187.21.7500-7510.2005
Multiple and interconnected pathways for L-lysine catabolism in Pseudomonas putida KT2440
Abstract
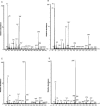
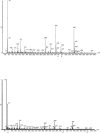
L-lysine catabolism in Pseudomonas putida KT2440 was generally thought to occur via the aminovalerate pathway. In this study we demonstrate the operation of the alternative aminoadipate pathway with the intermediates D-lysine, L-pipecolate, and aminoadipate. The simultaneous operation of both pathways for the use of L-lysine as the sole carbon and nitrogen source was confirmed genetically. Mutants with mutations in either pathway failed to use L-lysine as the sole carbon and nitrogen source, although they still used L-lysine as the nitrogen source, albeit at reduced growth rates. New genes were identified in both pathways, including the davB and davA genes that encode the enzymes involved in the oxidation of L-lysine to delta-aminovaleramide and the hydrolysis of the latter to delta-aminovalerate, respectively. The amaA, dkpA, and amaB genes, in contrast, encode proteins involved in the transformation of Delta1-piperidine-2-carboxylate into aminoadipate. Based on L-[U-13C, U-15N]lysine experiments, we quantified the relative use of pathways in the wild type and its isogenic mutants. The fate of 13C label of L-lysine indicates that in addition to the existing connection between the D- and L-lysine pathways at the early steps of the catabolism of L-lysine mediated by a lysine racemase, there is yet another interconnection at the lower end of the pathways in which aminoadipate is channeled to yield glutarate. This study establishes an unequivocal relationship between gene and pathway enzymes in the metabolism of L-lysine, which is of crucial importance for the successful colonization of the rhizosphere of plants by this microorganism.
Figures


Similar articles
-
D-lysine catabolic pathway in Pseudomonas putida: interrelations with L-lysine catabolism.J Bacteriol. 1974 Feb;117(2):753-64. doi: 10.1128/jb.117.2.753-764.1974. J Bacteriol. 1974. PMID: 4359655 Free PMC article.
-
Enzymatic production of 5-aminovalerate from L-lysine using L-lysine monooxygenase and 5-aminovaleramide amidohydrolase.Sci Rep. 2014 Jul 11;4:5657. doi: 10.1038/srep05657. Sci Rep. 2014. PMID: 25012259 Free PMC article.
-
Molecular characterization of lysR-lysXE, gcdR-gcdHG and amaR-amaAB operons for lysine export and catabolism: a comprehensive lysine catabolic network in Pseudomonas aeruginosa PAO1.Microbiology (Reading). 2016 May;162(5):876-888. doi: 10.1099/mic.0.000277. Epub 2016 Mar 11. Microbiology (Reading). 2016. PMID: 26967762
-
Identification of the initial steps in D-lysine catabolism in Pseudomonas putida.J Bacteriol. 2007 Apr;189(7):2787-92. doi: 10.1128/JB.01538-06. Epub 2007 Jan 26. J Bacteriol. 2007. PMID: 17259313 Free PMC article.
-
alpha-Aminoadipate pathway for the biosynthesis of lysine in lower eukaryotes.Crit Rev Microbiol. 1985;12(2):131-51. doi: 10.3109/10408418509104427. Crit Rev Microbiol. 1985. PMID: 3928261 Review.
Cited by
-
Evaluation of Heterologous Biosynthetic Pathways for Methanol-Based 5-Aminovalerate Production by Thermophilic Bacillus methanolicus.Front Bioeng Biotechnol. 2021 Jun 28;9:686319. doi: 10.3389/fbioe.2021.686319. eCollection 2021. Front Bioeng Biotechnol. 2021. PMID: 34262896 Free PMC article.
-
Regulation of Glutarate Catabolism by GntR Family Regulator CsiR and LysR Family Regulator GcdR in Pseudomonas putida KT2440.mBio. 2019 Jul 30;10(4):e01570-19. doi: 10.1128/mBio.01570-19. mBio. 2019. PMID: 31363033 Free PMC article.
-
Arginine as an environmental and metabolic cue for cyclic diguanylate signalling and biofilm formation in Pseudomonas putida.Sci Rep. 2020 Aug 12;10(1):13623. doi: 10.1038/s41598-020-70675-x. Sci Rep. 2020. PMID: 32788689 Free PMC article.
-
Genome-wide analysis of lysine catabolism in bacteria reveals new connections with osmotic stress resistance.ISME J. 2013 Dec;7(12):2400-10. doi: 10.1038/ismej.2013.123. Epub 2013 Jul 25. ISME J. 2013. PMID: 23887172 Free PMC article.
-
Overexpression of transport proteins improves the production of 5-aminovalerate from l-lysine in Escherichia coli.Sci Rep. 2016 Aug 11;6:30884. doi: 10.1038/srep30884. Sci Rep. 2016. PMID: 27510748 Free PMC article.
References
-
- Buell, C. R., V. Joardar, M. Lindeberg, J. Selengut, I. T. Paulsen, M. L. Gwinn, R. J. Dodson, R. T. Deboy, A. S. Durkin, J. F. Kolonay, R. Madupu, S. Daugherty, L. Brinkac, M. J. Beanan, D. H. Haft, W. C. Nelson, T. Davidsen, N. Zafar, L. Zhou, J. Lu, Q. Yuan, H. Khouri, N. Fedorova, B. Tran, D. Russell, K. Berry, T. Utterback, S. E. Van Aken, T. V. Feldblyum, M. D'Ascenzo, W. L. Deng, A. R. Ramos, J. R. Alfano, S. Cartinhour, A. K. Chatterjee, T. P. Delaney, S. G. Lazarowitz, G. B. Martin, D. J. Schneider, X. Tang, C. L. Bender, O. White, C. M. Fraser, and A. Collmer. 2003. The complete genome sequence of the Arabidopsis and tomato pathogen Pseudomonas syringae pv. tomato DC3000. Proc. Natl. Acad. Sci. USA 100:10181-10186. - PMC - PubMed
-
- Cao, X., J. Kolonay, Jr., K. A. Saxton, and R. A. Hartline. 1993. The OCT plasmid encodes D-lysine membrane transport and catabolic enzymes in Pseudomonas putida. Plasmid 30:83-89. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

