Molecular-level secondary structure, polymorphism, and dynamics of full-length alpha-synuclein fibrils studied by solid-state NMR
- PMID: 16247008
- PMCID: PMC1276071
- DOI: 10.1073/pnas.0506109102
Molecular-level secondary structure, polymorphism, and dynamics of full-length alpha-synuclein fibrils studied by solid-state NMR
Abstract
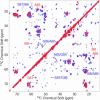
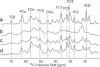
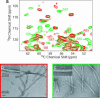
The 140-residue protein alpha-synuclein (AS) is able to form amyloid fibrils and as such is the main component of protein inclusions involved in Parkinson's disease. We have investigated the structure and dynamics of full-length AS fibrils by high-resolution solid-state NMR spectroscopy. Homonuclear and heteronuclear 2D and 3D spectra of fibrils grown from uniformly (13)C/(15)N-labeled AS and AS reverse-labeled for two of the most abundant amino acids, K and V, were analyzed. (13)C and (15)N signals exhibited linewidths of <0.7 ppm. Sequential assignments were obtained for 48 residues in the hydrophobic core region. We identified two different types of fibrils displaying chemical-shift differences of up to 13 ppm in the (15)N dimension and up to 5 ppm for backbone and side-chain (13)C chemical shifts. EM studies suggested that molecular structure is correlated with fibril morphology. Investigation of the secondary structure revealed that most amino acids of the core region belong to beta-strands with similar torsion angles in both conformations. Selection of regions with different mobility indicated the existence of monomers in the sample and allowed the identification of mobile segments of the protein within the fibril in the presence of monomeric protein. At least 35 C-terminal residues were mobile and lacked a defined secondary structure, whereas the N terminus was rigid starting from residue 22. Our findings agree well with the overall picture obtained with other methods and provide insight into the amyloid fibril structure and dynamics with residue-specific resolution.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

