Dual targeting is the rule for organellar aminoacyl-tRNA synthetases in Arabidopsis thaliana
- PMID: 16251277
- PMCID: PMC1283425
- DOI: 10.1073/pnas.0504682102
Dual targeting is the rule for organellar aminoacyl-tRNA synthetases in Arabidopsis thaliana
Abstract
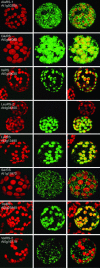
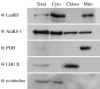
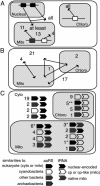
In plants, protein synthesis occurs in the cytosol, mitochondria, and plastids. Each compartment requires a full set of tRNAs and aminoacyl-tRNA synthetases. We have undertaken a systematic analysis of the targeting of organellar aminoacyl-tRNA synthetases in the model plant Arabidopsis thaliana. Dual targeting appeared to be a general rule. Among the 24 identified organellar aminoacyl-tRNA synthetases (aaRSs), 15 (and probably 17) are shared between mitochondria and plastids, and 5 are shared between cytosol and mitochondria (one of these aaRSs being present also in chloroplasts). Only two were shown to be uniquely chloroplastic and none to be uniquely mitochondrial. Moreover, there are no examples where the three aaRS genes originating from the three ancestral genomes still coexist. These results indicate that extensive exchange of aaRSs has occurred during evolution and that many are now shared between two or even three compartments. The findings have important implications for studies of the translation machinery in plants and on protein targeting and gene transfer in general.
Figures

Similar articles
-
Evolutionary history of Arabidopsis thaliana aminoacyl-tRNA synthetase dual-targeted proteins.Mol Biol Evol. 2011 Jan;28(1):79-85. doi: 10.1093/molbev/msq176. Epub 2010 Jul 12. Mol Biol Evol. 2011. PMID: 20624849
-
Defining the determinants for dual targeting of amino acyl-tRNA synthetases to mitochondria and chloroplasts.J Mol Biol. 2009 Nov 6;393(4):803-14. doi: 10.1016/j.jmb.2009.08.072. Epub 2009 Sep 4. J Mol Biol. 2009. PMID: 19733576
-
Requirement of aminoacyl-tRNA synthetases for gametogenesis and embryo development in Arabidopsis.Plant J. 2005 Dec;44(5):866-78. doi: 10.1111/j.1365-313X.2005.02580.x. Plant J. 2005. PMID: 16297076
-
Genomics and the evolution of aminoacyl-tRNA synthesis.Acta Biochim Pol. 2001;48(2):313-21. Acta Biochim Pol. 2001. PMID: 11732603 Review.
-
Nonconventional localizations of cytosolic aminoacyl-tRNA synthetases in yeast and human cells.Methods. 2017 Jan 15;113:91-104. doi: 10.1016/j.ymeth.2016.09.017. Epub 2016 Oct 7. Methods. 2017. PMID: 27725303 Review.
Cited by
-
An Enigmatic Stramenopile Sheds Light on Early Evolution in Ochrophyta Plastid Organellogenesis.Mol Biol Evol. 2022 Apr 11;39(4):msac065. doi: 10.1093/molbev/msac065. Mol Biol Evol. 2022. PMID: 35348760 Free PMC article.
-
The plant-specific ssDNA binding protein OSB1 is involved in the stoichiometric transmission of mitochondrial DNA in Arabidopsis.Plant Cell. 2006 Dec;18(12):3548-63. doi: 10.1105/tpc.106.042028. Epub 2006 Dec 22. Plant Cell. 2006. PMID: 17189341 Free PMC article.
-
The tomato chloroplast stromal proteome compendium elucidated by leveraging a plastid protein-localization prediction Atlas.Front Plant Sci. 2023 Aug 28;14:1020275. doi: 10.3389/fpls.2023.1020275. eCollection 2023. Front Plant Sci. 2023. PMID: 37701797 Free PMC article.
-
Two CRM protein subfamilies cooperate in the splicing of group IIB introns in chloroplasts.RNA. 2008 Nov;14(11):2319-32. doi: 10.1261/rna.1223708. Epub 2008 Sep 17. RNA. 2008. PMID: 18799595 Free PMC article.
-
Experimental analysis of the rice mitochondrial proteome, its biogenesis, and heterogeneity.Plant Physiol. 2009 Feb;149(2):719-34. doi: 10.1104/pp.108.131300. Epub 2008 Nov 14. Plant Physiol. 2009. PMID: 19010998 Free PMC article.
References
-
- Maréchal-Drouard, L., Weil, J. H. & Dietrich, A. (1993) Annu. Rev. Plant Physiol. Plant Mol. Biol. 44, 13-32.
-
- Duchêne, A. M. & Maréchal-Drouard, L. (2001) Biochem. Biophys. Res. Comm. 285, 1213-1216. - PubMed
-
- Akashi, K., Grandjean, O. & Small, I. (1998) FEBS Lett. 431, 39-44. - PubMed
-
- Duchêne, A. M., Peeters, N., Dietrich, A., Cosset, A., Small, I. D. & Wintz, H. (2001) J. Biol. Chem. 276, 15275-15283. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

