High potency of indolyl aryl sulfone nonnucleoside inhibitors towards drug-resistant human immunodeficiency virus type 1 reverse transcriptase mutants is due to selective targeting of different mechanistic forms of the enzyme
- PMID: 16251294
- PMCID: PMC1280147
- DOI: 10.1128/AAC.49.11.4546-4554.2005
High potency of indolyl aryl sulfone nonnucleoside inhibitors towards drug-resistant human immunodeficiency virus type 1 reverse transcriptase mutants is due to selective targeting of different mechanistic forms of the enzyme
Abstract
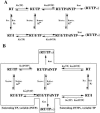
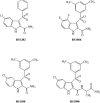
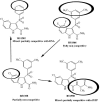
Indolyl aryl sulfone (IAS) nonnucleoside inhibitors have been shown to potently inhibit the growth of wild-type and drug-resistant human immunodeficiency virus type 1 (HIV-1), but their exact mechanism of action has not been elucidated yet. Here, we describe the mechanism of inhibition of HIV-1 reverse transcriptase (RT) by selected IAS derivatives. Our results showed that, depending on the substitutions introduced in the IAS common pharmacophore, these compounds can be made selective for different enzyme-substrate complexes. Moreover, we showed that the molecular basis for this selectivity was a different association rate of the drug to a particular enzymatic form along the reaction pathway. By comparing the activities of the different compounds against wild-type RT and the nonnucleoside reverse transcriptase inhibitor-resistant mutant Lys103Asn, it was possible to hypothesize, on the basis of their mechanism of action, a rationale for the design of drugs which could overcome the steric barrier imposed by the Lys103Asn mutation.
Figures

Similar articles
-
Non-nucleoside HIV-1 reverse transcriptase inhibitors di-halo-indolyl aryl sulfones achieve tight binding to drug-resistant mutants by targeting the enzyme-substrate complex.Antiviral Res. 2009 Jan;81(1):47-55. doi: 10.1016/j.antiviral.2008.09.008. Epub 2008 Nov 5. Antiviral Res. 2009. PMID: 18984007
-
Novel nonnucleoside inhibitors that select nucleoside inhibitor resistance mutations in human immunodeficiency virus type 1 reverse transcriptase.Antimicrob Agents Chemother. 2006 Aug;50(8):2772-81. doi: 10.1128/AAC.00127-06. Antimicrob Agents Chemother. 2006. PMID: 16870771 Free PMC article.
-
Mechanism of interaction of novel indolylarylsulfone derivatives with K103N and Y181I mutant HIV-1 reverse transcriptase in complex with its substrates.Antivir Chem Chemother. 2011 Nov 17;22(3):107-18. doi: 10.3851/IMP1855. Antivir Chem Chemother. 2011. PMID: 22095519
-
Mechanisms of resistance to nucleoside analogue inhibitors of HIV-1 reverse transcriptase.Virus Res. 2008 Jun;134(1-2):124-46. doi: 10.1016/j.virusres.2007.12.015. Epub 2008 Feb 12. Virus Res. 2008. PMID: 18272247 Review.
-
HIV-1 reverse transcriptase inhibitor resistance mutations and fitness: a view from the clinic and ex vivo.Virus Res. 2008 Jun;134(1-2):104-23. doi: 10.1016/j.virusres.2007.12.021. Epub 2008 Mar 4. Virus Res. 2008. PMID: 18289713 Review.
Cited by
-
Extracellular vesicles expressing a single-chain variable fragment of an HIV-1 specific antibody selectively target Env+ tissues.Theranostics. 2019 Jul 29;9(19):5657-5671. doi: 10.7150/thno.33925. eCollection 2019. Theranostics. 2019. PMID: 31534509 Free PMC article.
-
Status Presens of Antiviral Drugs And Strategies: Part I: DNA Viruses and Retroviruses.Adv Antivir Drug Des. 2007;5:1-58. doi: 10.1016/S1075-8593(06)05001-5. Epub 2007 Sep 2. Adv Antivir Drug Des. 2007. PMID: 32288472 Free PMC article. Review.
-
A new strategy to inhibit the excision reaction catalysed by HIV-1 reverse transcriptase: compounds that compete with the template-primer.Biochem J. 2007 Jul 1;405(1):165-71. doi: 10.1042/BJ20061831. Biochem J. 2007. PMID: 17355225 Free PMC article.
-
Biochemical mechanism of HIV-1 resistance to rilpivirine.J Biol Chem. 2012 Nov 2;287(45):38110-23. doi: 10.1074/jbc.M112.398180. Epub 2012 Sep 6. J Biol Chem. 2012. PMID: 22955279 Free PMC article.
-
Indolylarylsulfones, a fascinating story of highly potent human immunodeficiency virus type 1 non-nucleoside reverse transcriptase inhibitors.Antivir Chem Chemother. 2018 Jan-Dec;26:2040206617753443. doi: 10.1177/2040206617753443. Antivir Chem Chemother. 2018. PMID: 29417826 Free PMC article. Review.
References
-
- Bahar, I., B. Erman, R. L. Jernigan, A. R. Atilgan, and D. G. Covell. 1999. Collective motions in HIV-1 reverse transcriptase: examination of flexibility and enzyme function. J. Mol. Biol. 285:1023-1037. - PubMed
-
- Baldanti, F., S. Paolucci, G. Maga, N. Labo, U. Hubscher, A. Y. Skoblov, L. Victorova, S. Spadari, L. Minoli, and G. Gerna. 2003. Nevirapine-selected mutations Tyr181Ile/C of HIV-1 reverse transcriptase confer cross-resistance to stavudine. AIDS 17:1568-1570. - PubMed
-
- Basavapathruni, A., C. M. Bailey, and K. S. Anderson. 2004. Defining a molecular mechanism of synergy between nucleoside and nonnucleoside AIDS drugs. J. Biol. Chem. 279:6221-6224. (First published 13 January 2004.) - PubMed
-
- Blanca, G., F. Baldanti, S. Paolucci, A. Y. Skoblov, L. Victorova, U. Hubscher, G. Gerna, S. Spadari, and G. Maga. 2003. Nevirapine resistance mutation at codon 181 of the HIV-1 reverse transcriptase confers stavudine resistance by increasing nucleotide substrate discrimination and phosphorolytic activity. J. Biol. Chem. 278:15469-15472. (First published 24 February 2003.) - PubMed
-
- Buckheit, R. W., Jr., E. L. White, V. Fliakas-Boltz, J. Russell, T. L. Stup, T. L. Kinjerski, M. C. Osterling, A. Weigand, and J. P. Bader. 1999. Unique anti-human immunodeficiency virus activities of the nonnucleoside reverse transcriptase inhibitors calanolide A, costatolide, and dihydrocostatolide. Antimicrob. Agents Chemother. 43:1827-1834. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

