MAASE: an alternative splicing database designed for supporting splicing microarray applications
- PMID: 16251387
- PMCID: PMC1370865
- DOI: 10.1261/rna.2650905
MAASE: an alternative splicing database designed for supporting splicing microarray applications
Abstract
Alternative splicing is a prominent feature of higher eukaryotes. Understanding of the function of mRNA isoforms and the regulation of alternative splicing is a major challenge in the post-genomic era. The development of mRNA isoform sensitive microarrays, which requires precise splice-junction sequence information, is a promising approach. Despite the availability of a large number of mRNAs and ESTs in various databases and the efforts made to align transcript sequences to genomic sequences, existing alternative splicing databases do not offer adequate information in an appropriate format to aid in splicing array design. Here we describe our effort in constructing the Manually Annotated Alternatively Spliced Events (MAASE) database system, which is specifically designed to support splicing microarray applications. MAASE comprises two components: (1) a manual/computational annotation tool for the efficient extraction of critical sequence and functional information for alternative splicing events and (2) a user-friendly database of annotated events that allows convenient export of information to aid in microarray design and data analysis. We provide a detailed introduction and a step-by-step user guide to the MAASE database system to facilitate future large-scale annotation efforts, integration with other alternative splicing databases, and splicing array fabrication.
Figures



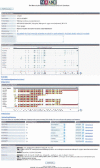


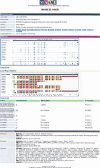
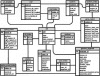
Similar articles
-
A database designed to computationally aid an experimental approach to alternative splicing.Pac Symp Biocomput. 2004:78-88. doi: 10.1142/9789812704856_0008. Pac Symp Biocomput. 2004. PMID: 14992494
-
Extent and diversity of human alternative splicing established by complementary database annotation and microarray analysis.OMICS. 2008 Mar;12(1):83-92. doi: 10.1089/omi.2007.0041. OMICS. 2008. PMID: 18266558
-
EuSplice: a unified resource for the analysis of splice signals and alternative splicing in eukaryotic genes.Bioinformatics. 2007 Jul 15;23(14):1815-23. doi: 10.1093/bioinformatics/btm084. Epub 2007 Mar 7. Bioinformatics. 2007. PMID: 17344236
-
Bioinformatics detection of alternative splicing.Methods Mol Biol. 2008;452:179-97. doi: 10.1007/978-1-60327-159-2_9. Methods Mol Biol. 2008. PMID: 18566765 Review.
-
Exploring the functional impact of alternative splicing on human protein isoforms using available annotation sources.Brief Bioinform. 2019 Sep 27;20(5):1754-1768. doi: 10.1093/bib/bby047. Brief Bioinform. 2019. PMID: 29931155 Free PMC article. Review.
Cited by
-
A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons.Genes Dev. 2007 Jul 1;21(13):1636-52. doi: 10.1101/gad.1558107. Genes Dev. 2007. PMID: 17606642 Free PMC article.
-
A procedure for identifying homologous alternative splicing events.BMC Bioinformatics. 2007 Jul 19;8:260. doi: 10.1186/1471-2105-8-260. BMC Bioinformatics. 2007. PMID: 17640387 Free PMC article.
-
Re-splicing of mature mRNA in cancer cells promotes activation of distant weak alternative splice sites.Nucleic Acids Res. 2012 Sep;40(16):7896-906. doi: 10.1093/nar/gks520. Epub 2012 Jun 6. Nucleic Acids Res. 2012. PMID: 22675076 Free PMC article.
-
Profiling alternatively spliced mRNA isoforms for prostate cancer classification.BMC Bioinformatics. 2006 Apr 11;7:202. doi: 10.1186/1471-2105-7-202. BMC Bioinformatics. 2006. PMID: 16608523 Free PMC article.
-
SpliceMiner: a high-throughput database implementation of the NCBI Evidence Viewer for microarray splice variant analysis.BMC Bioinformatics. 2007 Mar 5;8:75. doi: 10.1186/1471-2105-8-75. BMC Bioinformatics. 2007. PMID: 17338820 Free PMC article.
References
-
- Baelde, H.J., Eikmans, M., van Vliet, A.I., Bergijk, E.C., de Heer, E., and Bruijn, J.A. 2004. Alternatively spliced isoforms of fibronectin in immune-mediated glomerulosclerosis: The role of TGFβ and IL-4. J. Pathol. 204: 248–257. - PubMed
-
- Beck, S., Penque, D., Garcia, S., Gomes, A., Farinha, C., Mata, L., Gulbenkian, S., Gil Ferreira, K., Duarte, A., Pacheco, P., et al. 1999. Cystic fibrosis patients with the 3272–26A→G mutation have mild disease, leaky alternative mRNA splicing, and CFTR protein at the cell membrane. Hum. Mutat. 14: 133–144. - PubMed
-
- Caceres, J.F. and Kornblihtt, A.R. 2002. Alternative splicing: Multiple control mechanisms and involvement in human disease. Trends Genet. 18: 186–193. - PubMed
-
- Clark, T.A., Sugnet, C.W., and Ares Jr., M., 2002. Genome-wide analysis of mRNA processing in yeast using splicing-specific microarrays. Science 296: 907–910. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
