Dependence of the E. coli promoter strength and physical parameters upon the nucleotide sequence
- PMID: 16252339
- PMCID: PMC1390652
- DOI: 10.1631/jzus.2005.B1063
Dependence of the E. coli promoter strength and physical parameters upon the nucleotide sequence
Abstract
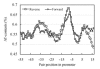
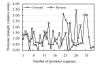
The energy of interaction between complementary nucleotides in promoter sequences of E. coli was calculated and visualized. The graphic method for presentation of energy properties of promoter sequences was elaborated on. Data obtained indicated that energy distribution through the length of promoter sequence results in picture with minima at -35, -8 and +7 regions corresponding to areas with elevated AT (adenine-thymine) content. The most important difference from the random sequences area is related to -8. Four promoter groups and their energy properties were revealed. The promoters with minimal and maximal energy of interaction between complementary nucleotides have low strengths, the strongest promoters correspond to promoter clusters characterized by intermediate energy values.
Figures



Similar articles
-
Regularities in the E. coli promoters composition in connection with the DNA strands interaction and promoter activity.J Zhejiang Univ Sci B. 2006 Dec;7(12):969-73. doi: 10.1631/jzus.2006.B0969. J Zhejiang Univ Sci B. 2006. PMID: 17111465 Free PMC article.
-
E. coli promoter prediction using feed-forward neural networks.Conf Proc IEEE Eng Med Biol Soc. 2006;2006:2025-7. doi: 10.1109/IEMBS.2006.260365. Conf Proc IEEE Eng Med Biol Soc. 2006. PMID: 17946085
-
The sequence upstream of the -10 consensus sequence modulates the strength and induction time of stationary-phase promoters in Escherichia coli.Appl Microbiol Biotechnol. 2005 Dec;69(3):312-20. doi: 10.1007/s00253-005-0016-8. Epub 2005 Nov 15. Appl Microbiol Biotechnol. 2005. PMID: 16088348
-
[Specificiety of DNA-protein interactions within transcription complexes of Escherichia coli].Mol Biol (Mosk). 2004 Sep-Oct;38(5):786-97. Mol Biol (Mosk). 2004. PMID: 15554182 Review. Russian.
-
Computational analyses of eukaryotic promoters.BMC Bioinformatics. 2007 Sep 27;8 Suppl 6(Suppl 6):S3. doi: 10.1186/1471-2105-8-S6-S3. BMC Bioinformatics. 2007. PMID: 17903284 Free PMC article. Review.
Cited by
-
Regularities in the E. coli promoters composition in connection with the DNA strands interaction and promoter activity.J Zhejiang Univ Sci B. 2006 Dec;7(12):969-73. doi: 10.1631/jzus.2006.B0969. J Zhejiang Univ Sci B. 2006. PMID: 17111465 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

