Kinetic analyses of the surface-transmembrane disulfide bond isomerization-controlled fusion activation pathway in Moloney murine leukemia virus
- PMID: 16254321
- PMCID: PMC1280236
- DOI: 10.1128/JVI.79.22.13856-13864.2005
Kinetic analyses of the surface-transmembrane disulfide bond isomerization-controlled fusion activation pathway in Moloney murine leukemia virus
Abstract
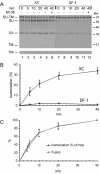
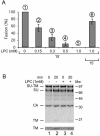
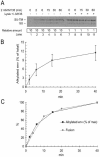
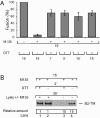
The surface (SU) and transmembrane (TM) subunits of Moloney murine leukemia virus (Mo-MLV) Env are disulfide linked. The linking cysteine in SU is part of a conserved CXXC motif in which the other cysteine carries a free thiol. Recently, we showed that receptor binding activates its free thiol to isomerize the intersubunit disulfide bond into a disulfide within the motif instead (M. Wallin, M. Ekström and H. Garoff, EMBO J. 23:54-65, 2004). This facilitated SU dissociation and activation of TM for membrane fusion. The evidence was mainly based on the finding that alkylation of the CXXC-thiol prevented isomerization. This arrested membrane fusion, but the activity could be rescued by cleaving the intersubunit disulfide bond with dithiothreitol (DTT). Here, we demonstrate directly that receptor binding causes SU-TM disulfide bond isomerization in a subfraction of the viral Envs. The kinetics of the isomerization followed that of virus-cell membrane fusion. Arresting the fusion with lysophosphatidylcholine did not arrest isomerization, suggesting that isomerization precedes the hemifusion stage of fusion. Our earlier finding that native Env was not possible to alkylate but required isomerization induction by receptor binding intimated that alkylation trapped an intermediate form of Env. To further clarify this possibility, we analyzed the kinetics by which the alkylation-sensitive Env was generated during fusion. We found that it followed the fusion kinetics. In contrast, the release of fusion from alkylated, isomerization-blocked virus by DTT reduction of the SU-TM disulfide bond was much faster. These results suggest that the alkylation-sensitive form of Env is a true intermediate in the fusion activation pathway of Env.
Figures
Similar articles
-
The fusion-controlling disulfide bond isomerase in retrovirus Env is triggered by protein destabilization.J Virol. 2005 Feb;79(3):1678-85. doi: 10.1128/JVI.79.3.1678-1685.2005. J Virol. 2005. PMID: 15650193 Free PMC article.
-
The conserved His8 of the Moloney murine leukemia virus Env SU subunit directs the activity of the SU-TM disulphide bond isomerase.Virology. 2007 Apr 25;361(1):149-60. doi: 10.1016/j.virol.2006.11.013. Epub 2006 Dec 19. Virology. 2007. PMID: 17182074
-
Isomerization of the intersubunit disulphide-bond in Env controls retrovirus fusion.EMBO J. 2004 Jan 14;23(1):54-65. doi: 10.1038/sj.emboj.7600012. Epub 2003 Dec 11. EMBO J. 2004. PMID: 14685283 Free PMC article.
-
[Chemokine receptors and its importance in the replication cycle of human immunodeficiency virus: clinical and therapeutic implications].Acta Med Port. 2008 Sep-Oct;21(5):497-504. Epub 2009 Jan 16. Acta Med Port. 2008. PMID: 19187693 Review. Portuguese.
-
Protein disulfides and protein disulfide oxidoreductases in hyperthermophiles.FEBS J. 2006 Sep;273(18):4170-85. doi: 10.1111/j.1742-4658.2006.05421.x. Epub 2006 Aug 23. FEBS J. 2006. PMID: 16930136 Review.
Cited by
-
Role of thiol/disulfide exchange in newcastle disease virus entry.J Virol. 2009 Jan;83(1):241-9. doi: 10.1128/JVI.01407-08. Epub 2008 Oct 15. J Virol. 2009. PMID: 18922867 Free PMC article.
-
Stabilization of TM trimer interactions during activation of moloney murine leukemia virus Env.J Virol. 2008 Mar;82(5):2358-66. doi: 10.1128/JVI.01931-07. Epub 2007 Dec 19. J Virol. 2008. PMID: 18094169 Free PMC article.
-
Receptor-triggered but alkylation-arrested env of murine leukemia virus reveals the transmembrane subunit in a prehairpin conformation.J Virol. 2006 Oct;80(19):9921-5. doi: 10.1128/JVI.00380-06. J Virol. 2006. PMID: 16973599 Free PMC article.
-
Structures and mechanisms of viral membrane fusion proteins: multiple variations on a common theme.Crit Rev Biochem Mol Biol. 2008 May-Jun;43(3):189-219. doi: 10.1080/10409230802058320. Crit Rev Biochem Mol Biol. 2008. PMID: 18568847 Free PMC article. Review.
-
Cell-type specific requirements for thiol/disulfide exchange during HIV-1 entry and infection.Retrovirology. 2012 Dec 3;9:97. doi: 10.1186/1742-4690-9-97. Retrovirology. 2012. PMID: 23206338 Free PMC article.
References
-
- Bullough, P. A., F. M. Hughson, J. J. Skehel, and D. C. Wiley. 1994. Structure of influenza haemagglutinin at the pH of membrane fusion. Nature 371:37-43. - PubMed
-
- Chan, D. C., D. Fass, J. M. Berger, and P. S. Kim. 1997. Core structure of gp41 from the HIV envelope glycoprotein. Cell 89:263-273. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

