An essential RNase III insertion editing endonuclease in Trypanosoma brucei
- PMID: 16269544
- PMCID: PMC1283813
- DOI: 10.1073/pnas.0506133102
An essential RNase III insertion editing endonuclease in Trypanosoma brucei
Abstract
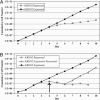
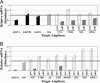
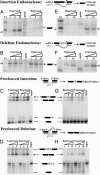
RNA editing adds and deletes uridine nucleotides in many preedited mRNAs to create translatable mRNAs in the mitochondria of the parasite Trypanosoma brucei. Kinetoplastid RNA editing protein B3 (KREPB3, formerly TbMP61) is part of the multiprotein complex that catalyzes editing in T. brucei and contains an RNase III motif that suggests nuclease function. Repression of KREPB3 expression, either by RNA interference in procyclic forms (PFs) or by conditional inactivation of an ectopic KREPB3 allele in bloodstream forms (BFs) that lack both endogenous alleles, strongly inhibited growth and in vivo editing in PFs and completely blocked them in BFs. KREPB3 repression inhibited cleavage of insertion editing substrates but not deletion editing substrates in vitro, whereas the terminal uridylyl transferase, U-specific exoribonuclease, and ligase activities of editing were unaffected, and approximately 20S editosomes were retained. Expression of KREPB3 alleles with single amino acid mutations in the RNase III motif had similar consequences. These data indicate that KREPB3 is an RNA editing endonuclease that is specific for insertion sites and is accordingly renamed KREN2 (kinetoplastid RNA editing endonuclease 2).
Figures


Similar articles
-
A deletion site editing endonuclease in Trypanosoma brucei.Mol Cell. 2005 Nov 11;20(3):403-12. doi: 10.1016/j.molcel.2005.09.016. Mol Cell. 2005. PMID: 16285922
-
Compositionally and functionally distinct editosomes in Trypanosoma brucei.RNA. 2006 Jun;12(6):1038-49. doi: 10.1261/rna.45506. Epub 2006 Apr 12. RNA. 2006. PMID: 16611942 Free PMC article.
-
Editosome RNase III domain interactions are essential for editing and differ between life cycle stages in Trypanosoma brucei.RNA. 2019 Sep;25(9):1150-1163. doi: 10.1261/rna.071258.119. Epub 2019 Jun 6. RNA. 2019. PMID: 31171708 Free PMC article.
-
U-Insertion/Deletion mRNA-Editing Holoenzyme: Definition in Sight.Trends Parasitol. 2016 Feb;32(2):144-156. doi: 10.1016/j.pt.2015.10.004. Epub 2015 Nov 10. Trends Parasitol. 2016. PMID: 26572691 Free PMC article. Review.
-
'Gestalt,' composition and function of the Trypanosoma brucei editosome.Annu Rev Microbiol. 2012;66:65-82. doi: 10.1146/annurev-micro-092611-150150. Annu Rev Microbiol. 2012. PMID: 22994488 Review.
Cited by
-
Structures of the T. brucei kRNA editing factor MRB1590 reveal unique RNA-binding pore motif contained within an ABC-ATPase fold.Nucleic Acids Res. 2015 Aug 18;43(14):7096-109. doi: 10.1093/nar/gkv647. Epub 2015 Jun 27. Nucleic Acids Res. 2015. PMID: 26117548 Free PMC article.
-
Trypanosoma brucei Tb927.2.6100 is an essential protein associated with kinetoplast DNA.Eukaryot Cell. 2013 Jul;12(7):970-8. doi: 10.1128/EC.00352-12. Epub 2013 May 6. Eukaryot Cell. 2013. PMID: 23650088 Free PMC article.
-
Trypanosome Letm1 protein is essential for mitochondrial potassium homeostasis.J Biol Chem. 2013 Sep 13;288(37):26914-25. doi: 10.1074/jbc.M113.495119. Epub 2013 Jul 26. J Biol Chem. 2013. PMID: 23893410 Free PMC article.
-
Structural basis for UTP specificity of RNA editing TUTases from Trypanosoma brucei.EMBO J. 2005 Dec 7;24(23):4007-17. doi: 10.1038/sj.emboj.7600861. Epub 2005 Nov 10. EMBO J. 2005. PMID: 16281058 Free PMC article.
-
Distinct and overlapping functions of MRP1/2 and RBP16 in mitochondrial RNA metabolism.Mol Cell Biol. 2009 Oct;29(19):5214-25. doi: 10.1128/MCB.00520-09. Epub 2009 Jul 20. Mol Cell Biol. 2009. PMID: 19620277 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

