Identification and characterization of a novel biotin biosynthesis gene in Saccharomyces cerevisiae
- PMID: 16269718
- PMCID: PMC1287709
- DOI: 10.1128/AEM.71.11.6845-6855.2005
Identification and characterization of a novel biotin biosynthesis gene in Saccharomyces cerevisiae
Abstract
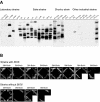
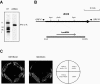
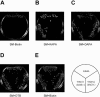
Yeast Saccharomyces cerevisiae cells generally cannot synthesize biotin, a vitamin required for many carboxylation reactions. Although sake yeasts, which are used for Japanese sake brewing, are classified as S. cerevisiae, they do not require biotin for their growth. In this study, we identified a novel open reading frame (ORF) in the genome of one strain of sake yeast that we speculated to be involved in biotin synthesis. Homologs of this gene are widely distributed in the genomes of sake yeasts. However, they are not found in many laboratory strains and strains used for wine making and beer brewing. This ORF was named BIO6 because it has 52% identity with BIO3, a biotin biosynthesis gene of a laboratory strain. Further research showed that yeasts without the BIO6 gene are auxotrophic for biotin, whereas yeasts holding the BIO6 gene are prototrophic for biotin. The BIO6 gene was disrupted in strain A364A, which is a laboratory strain with one copy of the BIO6 gene. Although strain A364A is prototrophic for biotin, a BIO6 disrupted mutant was found to be auxotrophic for biotin. The BIO6 disruptant was able to grow in biotin-deficient medium supplemented with 7-keto-8-amino-pelargonic acid (KAPA), while the bio3 disruptant was not able to grow in this medium. These results suggest that Bio6p acts in an unknown step of biotin synthesis before KAPA synthesis. Furthermore, we demonstrated that expression of the BIO6 gene, like that of other biotin synthesis genes, was upregulated by depletion of biotin. We conclude that the BIO6 gene is a novel biotin biosynthesis gene of S. cerevisiae.
Figures

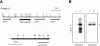


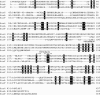

References
-
- Adams, A., D. E. Gottschling, C. Kaiser, and T. Stearns. 1997. Methods in yeast genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
-
- Baldet, P., and M. L. Ruffet. 1996. Biotin synthesis in higher plants: isolation of a cDNA encoding Arabidopsis thaliana bioB-gene product equivalent by functional complementation of a biotin auxotroph mutant bioB105 of Escherichia coli K12. C. R. Acad. Sci. Ser. III 319:99-106. - PubMed
-
- Becker, D. M., and L. Guarente. 1991. High-efficiency transformation of yeast by electroporation. Methods Enzymol. 194:182-187. - PubMed
-
- Betz, H., H. Hinze, and H. Holzer. 1974. Isolation and properties of two inhibitors of proteinase B from yeast. J. Biol. Chem. 249:4515-4521. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

