Catch-bond model derived from allostery explains force-activated bacterial adhesion
- PMID: 16272438
- PMCID: PMC1367101
- DOI: 10.1529/biophysj.105.066548
Catch-bond model derived from allostery explains force-activated bacterial adhesion
Abstract
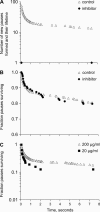
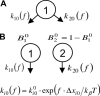
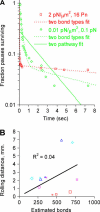
High shear enhances the adhesion of Escherichia coli bacteria binding to mannose coated surfaces via the adhesin FimH, raising the question as to whether FimH forms catch bonds that are stronger under tensile mechanical force. Here, we study the length of time that E. coli pause on mannosylated surfaces and report a double exponential decay in the duration of the pauses. This double exponential decay is unlike previous single molecule or whole cell data for other catch bonds, and indicates the existence of two distinct conformational states. We present a mathematical model, derived from the common notion of chemical allostery, which describes the lifetime of a catch bond in which mechanical force regulates the transitions between two conformational states that have different unbinding rates. The model explains these characteristics of the data: a double exponential decay, an increase in both the likelihood and lifetime of the high-binding state with shear stress, and a biphasic effect of force on detachment rates. The model parameters estimated from the data are consistent with the force-induced structural changes shown earlier in FimH. This strongly suggests that FimH forms allosteric catch bonds. The model advances our understanding of both catch bonds and the role of allostery in regulating protein activity.
Figures





References
-
- Thomas, W. E., E. Trintchina, M. Forero, V. Vogel, and E. V. Sokurenko. 2002. Bacterial adhesion to target cells enhanced by shear force. Cell. 109:913–923. - PubMed
-
- Isberg, R. R., and P. Barnes. 2002. Dancing with the host; flow-dependent bacterial adhesion. Cell. 110:1–4. - PubMed
-
- Konstantopoulos, K., W. D. Hanley, and D. Wirtz. 2003. Receptor-ligand binding: ‘catch’ bonds finally caught. Curr. Biol. 13:R611–R613. - PubMed
-
- Thomas, W. E., L. Nilsson, M. Forero, E. V. Sokurenko, and V. Vogel. 2004. ‘Stick-and-roll’ bacterial adhesion mediated by catch-bonds. Mol. Microbiol. 53:1545–1557. - PubMed
-
- Finger, E. B., K. D. Puri, R. Alon, M. B. Lawrence, U. H. von Andrian, and T. A. Springer. 1996. Adhesion through L-selectin requires a threshold hydrodynamic shear. Nature. 379:266–269. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

