Population structure and properties of Candida albicans, as determined by multilocus sequence typing
- PMID: 16272493
- PMCID: PMC1287804
- DOI: 10.1128/JCM.43.11.5601-5613.2005
Population structure and properties of Candida albicans, as determined by multilocus sequence typing
Abstract
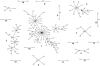
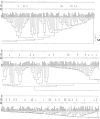
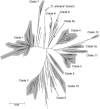
We submitted a panel of 416 isolates of Candida albicans from separate sources to multilocus sequence typing (MLST). The data generated determined a population structure in which four major clades of closely related isolates were delineated, together with eight minor clades comprising five or more isolates. By Fisher's exact test, a statistically significant association was found between particular clades and the anatomical source, geographical source, ABC genotype, decade of isolation, and homozygosity versus heterozygosity at the mating type-like locus (MTL) of the isolates in the clade. However, these associations may have been influenced by confounding variables, since in a univariate analysis of variance, only the clade associations with ABC type and anatomical source emerged as statistically significant, providing the first indication of possible differences between C. albicans strain type clades and their propensity to infect or colonize different anatomical locations. There were no significant differences between clades with respect to distributions of isolates resistant to fluconazole, itraconazole, or flucytosine. However, the majority of flucytosine-resistant isolates belonged to clade 1, and these isolates, but not flucytosine-resistant isolates in other clades, bore a unique mutation in the FUR1 gene that probably accounts for their resistance. A significantly higher proportion of isolates resistant to fluconazole, itraconazole, and flucytosine were homozygous at the MTL, suggesting that antifungal pressure may trigger a common mechanism that leads both to resistance and to MTL homozygosity. The utility of MLST for determining clade assignments of clinical isolates will form the basis for strain selection for future research into C. albicans virulence.
Figures

Similar articles
-
In Candida albicans, resistance to flucytosine and terbinafine is linked to MAT locus homozygosity and multilocus sequence typing clade 1.FEMS Yeast Res. 2009 Oct;9(7):1091-101. doi: 10.1111/j.1567-1364.2009.00577.x. Epub 2009 Sep 7. FEMS Yeast Res. 2009. PMID: 19799637
-
Molecular epidemiology of invasive Candida albicans at a tertiary hospital in northern Taiwan from 2003 to 2011.Med Mycol. 2015 Nov;53(8):828-36. doi: 10.1093/mmy/myv065. Epub 2015 Sep 1. Med Mycol. 2015. PMID: 26333357
-
Clade-related amphotericin B resistance among South African Candida albicans isolates.Diagn Microbiol Infect Dis. 2005 Sep;53(1):29-31. doi: 10.1016/j.diagmicrobio.2005.03.013. Diagn Microbiol Infect Dis. 2005. PMID: 16182076
-
Molecular phylogenetics and epidemiology of Candida albicans.Future Microbiol. 2010 Jan;5(1):67-79. doi: 10.2217/fmb.09.113. Future Microbiol. 2010. PMID: 20020830 Review.
-
Molecular epidemiology, phylogeny and evolution of Candida albicans.Infect Genet Evol. 2014 Jan;21:166-78. doi: 10.1016/j.meegid.2013.11.008. Epub 2013 Nov 19. Infect Genet Evol. 2014. PMID: 24269341 Review.
Cited by
-
Molecular fingerprints to identify Candida species.Biomed Res Int. 2013;2013:923742. doi: 10.1155/2013/923742. Epub 2013 Jun 17. Biomed Res Int. 2013. PMID: 23844370 Free PMC article. Review.
-
Alternative mating type configurations (a/α versus a/a or α/α) of Candida albicans result in alternative biofilms regulated by different pathways.PLoS Biol. 2011 Aug;9(8):e1001117. doi: 10.1371/journal.pbio.1001117. Epub 2011 Aug 2. PLoS Biol. 2011. PMID: 21829325 Free PMC article.
-
Phenotypic screening, transcriptional profiling, and comparative genomic analysis of an invasive and non-invasive strain of Candida albicans.BMC Microbiol. 2008 Oct 24;8:187. doi: 10.1186/1471-2180-8-187. BMC Microbiol. 2008. PMID: 18950481 Free PMC article.
-
Fluconazole Resistance Patterns in Candida Species that Colonize Women with HIV Infection.Curr Ther Res Clin Exp. 2014 Sep 28;76:84-9. doi: 10.1016/j.curtheres.2014.07.002. eCollection 2014 Dec. Curr Ther Res Clin Exp. 2014. PMID: 25352939 Free PMC article.
-
Optimized multilocus sequence typing (MLST) scheme for Trypanosoma cruzi.PLoS Negl Trop Dis. 2014 Aug 28;8(8):e3117. doi: 10.1371/journal.pntd.0003117. eCollection 2014 Aug. PLoS Negl Trop Dis. 2014. PMID: 25167160 Free PMC article.
References
-
- Blignaut, E., C. Pujol, S. Lockhart, S. Joly, and D. R. Soll. 2002. A new clade of Candida albicans among South African oral yeast isolates. J. Dent. Res. 81:2231.
-
- Bougnoux, M.-E., A. D. M., S. Morand, M. Théraud, B. G. Spratt, and C. d'Enfert. 2004. Multilocus sequence typing of Candida albicans: strategies, data exchange and applications. Infect. Genet. Evol. 4:243-252. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

