RNA-dependent integrin alpha3 protein localization regulated by the Muscleblind-like protein MLP1
- PMID: 16273094
- PMCID: PMC2365307
- DOI: 10.1038/ncb1335
RNA-dependent integrin alpha3 protein localization regulated by the Muscleblind-like protein MLP1
Erratum in
- Nat Cell Biol. 2006 Jan;8(1):100
Abstract
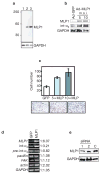
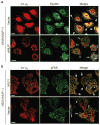
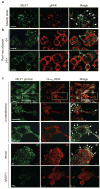
We show that localized expression of the integrin alpha3 protein is regulated at the level of RNA localization by the human homologue of Drosophila Muscleblind, MLP1/MBLL/MBNL2, a unique Cys3His zinc-finger protein. This is supported by the following observations: MLP1 knockdown abolishes localization of integrin alpha3 to the adhesion complexes; MLP1 is localized in adhesion plaques that contain phospho-focal adhesion kinase; this localization is microtubule-dependent; integrin alpha3 transcripts colocalize with MLP1 in distinct cytoplasmic loci; integrin alpha3 transcripts are physically associated with MLP1 in cells and MLP1 binds to a specific ACACCC motif in the integrin alpha3 3' untranslated region (UTR) in vitro; and a green fluorescent protein (GFP) open reading frame-integrin alpha3 3' UTR chimeric gene directs GFP protein localization to distinct cytoplasmic loci near the cell periphery, which is dependent on MLP1 and is mediated by the ACACCC motif but is independent of the integrin alpha3 signal peptide.
Figures


References
-
- Begemann G, Paricio N, Artero R, Kiss I, Perez-Alonso M, Mlodzik M. Muscleblind, a gene required for photoreceptor differentiation in Drosophila, encodes novel nuclear Cys3His-type zinc-finger-containing proteins. Development. 1997;124:4321–4331. - PubMed
-
- Artero R, et al. The muscleblind gene participates in the organization of Z-bands and epidermal attachments of Drosophila muscles and is regulated by Dmef2. Dev Biol. 1998;195:131–143. - PubMed
-
- Fardaei M, et al. Three proteins, MBNL, MBLL and MBXL, co-localize in vivo with nuclear foci of expanded-repeat transcripts in DM1 and DM2 cells. Hum Mol Genet. 2002;11:805–814. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

