Molecular characterization of adeno-associated viruses infecting children
- PMID: 16282478
- PMCID: PMC1287571
- DOI: 10.1128/JVI.79.23.14781-14792.2005
Molecular characterization of adeno-associated viruses infecting children
Abstract
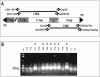
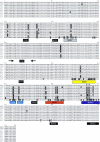
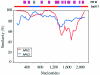
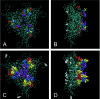
Although adeno-associated virus (AAV) infection is common in humans, the biology of natural infection is poorly understood. Since it is likely that many primary AAV infections occur during childhood, we set out to characterize the frequency and complexity of circulating AAV isolates in fresh and archived frozen human pediatric tissues. Total cellular DNA was isolated from 175 tissue samples including freshly collected tonsils (n = 101) and archived frozen samples representing spleen (n = 21), lung (n = 16), muscle (n = 15), liver (n = 19), and heart (n = 3). Samples were screened for the presence of AAV and adenovirus sequences by PCR using degenerate primers. AAV DNA was detected in 7 of 101 (7%) tonsil samples and two of 74 other tissues (one spleen and one lung). Adenovirus sequences were identified in 19 of 101 tonsils (19%), but not in any other tissues. Complete capsid gene sequences were recovered from all nine AAV-positive tissues. Sequence analyses showed that eight of the capsid sequences were AAV2-like (approximately 98% amino acid identity), while the single spleen isolate was intermediate between serotypes 2 and 3. Comparison to the available AAV2 crystal structure revealed that the majority of the amino acid substitutions mapped to surface-exposed hypervariable domains. To further characterize the AAV capsid structure in these samples, we used a novel linear rolling-circle amplification method to amplify episomal AAV DNA and isolate infectious molecular clones from several human tissues. Serotype 2-like viruses were generated from these DNA clones and interestingly, failed to bind to a heparin sulfate column. Inspection of the capsid sequence from these two clones (and the other six AAV2-like isolates) revealed that they lacked arginine residues at positions 585 and 588 of the capsid protein, which are thought to be essential for interaction with the heparin sulfate proteoglycan coreceptor. These data provide a framework with which to explore wild-type AAV persistence in vivo and provide additional tools to further define the biodistribution and form of AAV in human tissues.
Figures



References
-
- Bantel-Schaal, U., and H. zur Hausen. 1984. Characterization of the DNA of a defective human parvovirus isolated from a genital site. Virology 134:52-63. - PubMed
-
- Blacklow, N. R., M. D. Hoggan, A. Z. Kapikian, J. B. Austin, and W. P. Rowe. 1968. Epidemiology of adenovirus-associated virus infection in a nursery population. Am. J. Epidemiol. 88:368-378. - PubMed
-
- Blacklow, N. R., M. D. Hoggan, and W. P. Rowe. 1968. Serologic evidence for human infection with adenovirus-associated viruses. J. Natl. Cancer Inst. 40:319-327. - PubMed
-
- Blacklow, N. R., M. D. Hoggan, M. S. Sereno, C. D. Brandt, H. W. Kim, R. H. Parrott, and R. M. Chanock. 1971. A seroepidemiologic study of adenovirus-associated virus infection in infants and children. Am. J. Epidemiol. 94:359-366. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

