The hematopoietic transcription factor AML1 (RUNX1) is negatively regulated by the cell cycle protein cyclin D3
- PMID: 16287839
- PMCID: PMC1291252
- DOI: 10.1128/MCB.25.23.10205-10219.2005
The hematopoietic transcription factor AML1 (RUNX1) is negatively regulated by the cell cycle protein cyclin D3
Abstract
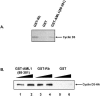
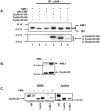
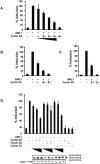
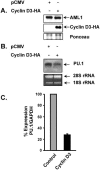
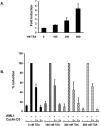
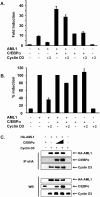
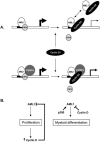
The family of cyclin D proteins plays a crucial role in the early events of the mammalian cell cycle. Recent studies have revealed the involvement of AML1 transactivation activity in promoting cell cycle progression through the induction of cyclin D proteins. This information in combination with our previous observation that a region in AML1 between amino acids 213 and 289 is important for its function led us to investigate prospective proteins associating with this region. We identified cyclin D3 by a yeast two-hybrid screen and detected AML1 interaction with the cyclin D family by both in vitro pull-down and in vivo coimmunoprecipitation assays. Furthermore, we demonstrate that cyclin D3 negatively regulates the transactivation activity of AML1 in a dose-dependent manner by competing with CBFbeta for AML1 association, leading to a decreased binding affinity of AML1 for its target DNA sequence. AML1 and its fusion protein AML1-ETO have been shown to shorten and prolong the mammalian cell cycle, respectively. In addition, AML1 promotes myeloid cell differentiation. Thus, our observations suggest that the direct association of cyclin D3 with AML1 functions as a putative feedback mechanism to regulate cell cycle progression and differentiation.
Figures



Similar articles
-
AML1/RUNX1 increases during G1 to S cell cycle progression independent of cytokine-dependent phosphorylation and induces cyclin D3 gene expression.J Biol Chem. 2004 Apr 9;279(15):15678-87. doi: 10.1074/jbc.M310023200. Epub 2004 Jan 27. J Biol Chem. 2004. PMID: 14747476
-
AML1 stimulates G1 to S progression via its transactivation domain.Oncogene. 2002 May 9;21(20):3247-52. doi: 10.1038/sj.onc.1205447. Oncogene. 2002. PMID: 12082641
-
AML1/RUNX1 phosphorylation by cyclin-dependent kinases regulates the degradation of AML1/RUNX1 by the anaphase-promoting complex.Mol Cell Biol. 2006 Oct;26(20):7420-9. doi: 10.1128/MCB.00597-06. Epub 2006 Aug 5. Mol Cell Biol. 2006. PMID: 17015473 Free PMC article.
-
AML1/Runx1 as a versatile regulator of hematopoiesis: regulation of its function and a role in adult hematopoiesis.Int J Hematol. 2006 Aug;84(2):136-42. doi: 10.1532/IJH97.06070. Int J Hematol. 2006. PMID: 16926135 Review.
-
Post-translational modifications of Runx1 regulate its activity in the cell.Blood Cells Mol Dis. 2009 Jul-Aug;43(1):30-4. doi: 10.1016/j.bcmd.2009.03.005. Epub 2009 Apr 21. Blood Cells Mol Dis. 2009. PMID: 19386523 Free PMC article. Review.
Cited by
-
CBFB-MYH11 hinders early T-cell development and induces massive cell death in the thymus.Blood. 2007 Apr 15;109(8):3432-40. doi: 10.1182/blood-2006-10-051508. Epub 2006 Dec 21. Blood. 2007. PMID: 17185462 Free PMC article.
-
Essential role for cyclin D3 in granulocyte colony-stimulating factor-driven expansion of neutrophil granulocytes.Mol Cell Biol. 2006 Nov;26(21):8052-60. doi: 10.1128/MCB.00800-06. Epub 2006 Sep 5. Mol Cell Biol. 2006. PMID: 16954383 Free PMC article.
-
Transcriptional Regulatory Networks in Hepatitis C Virus-induced Hepatocellular Carcinoma.Sci Rep. 2018 Sep 24;8(1):14234. doi: 10.1038/s41598-018-32464-5. Sci Rep. 2018. PMID: 30250040 Free PMC article.
-
Transcription factors Runx1 to 3 are expressed in the lacrimal gland epithelium and are involved in regulation of gland morphogenesis and regeneration.Invest Ophthalmol Vis Sci. 2013 May 1;54(5):3115-25. doi: 10.1167/iovs.13-11791. Invest Ophthalmol Vis Sci. 2013. PMID: 23532528 Free PMC article.
-
Cyclin-dependent kinase phosphorylation of RUNX1/AML1 on 3 sites increases transactivation potency and stimulates cell proliferation.Blood. 2008 Feb 1;111(3):1193-200. doi: 10.1182/blood-2007-08-109702. Epub 2007 Nov 14. Blood. 2008. PMID: 18003885 Free PMC article.
References
-
- Bates, S., and G. Peters. 1995. Cyclin D1 as a cellular proto-oncogene. Semin. Cancer Biol. 6:73-82. - PubMed
-
- Behre, G., P. Zhang, D. E. Zhang, and D. G. Tenen. 1999. Analysis of the modulation of transcriptional activity in myelopoiesis and leukemogenesis. Methods 17:231-237. - PubMed
-
- Bernardin, F., and A. D. Friedman. 2002. AML1 stimulates G1 to S progression via its transactivation domain. Oncogene 21:3247-3252. - PubMed
-
- Bernardin-Fried, F., T. Kummalue, S. Leijen, M. I. Collector, K. Ravid, and A. D. Friedman. 2004. AML1/RUNX1 increases during G1 to S cell cycle progression independent of cytokine-dependent phosphorylation and induces cyclin D3 gene expression. J. Biol. Chem. 279:15678-15687. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
