Histone-modifying complexes regulate gene expression pertinent to the differentiation of the protozoan parasite Toxoplasma gondii
- PMID: 16287846
- PMCID: PMC1291236
- DOI: 10.1128/MCB.25.23.10301-10314.2005
Histone-modifying complexes regulate gene expression pertinent to the differentiation of the protozoan parasite Toxoplasma gondii
Abstract
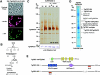
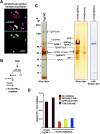
Pathogenic apicomplexan parasites like Toxoplasma and Plasmodium (malaria) have complex life cycles consisting of multiple stages. The ability to differentiate from one stage to another requires dramatic transcriptional changes, yet there is a paucity of transcription factors in these protozoa. In contrast, we show here that Toxoplasma possesses extensive chromatin remodeling machinery that modulates gene expression relevant to differentiation. We find that, as in other eukaryotes, histone acetylation and arginine methylation are marks of gene activation in Toxoplasma. We have identified mediators of these histone modifications, as well as a histone deacetylase (HDAC), and correlate their presence at target promoters in a stage-specific manner. We purified the first HDAC complex from apicomplexans, which contains novel components in addition to others previously reported in eukaryotes. A Toxoplasma orthologue of the arginine methyltransferase CARM1 appears to work in concert with the acetylase TgGCN5, which exhibits an unusual bias for H3 [K18] in vitro. Inhibition of TgCARM1 induces differentiation, showing that the parasite life cycle can be manipulated by interfering with epigenetic machinery. This may lead to new approaches for therapy against protozoal diseases and highlights Toxoplasma as an informative model to study the evolution of epigenetics in eukaryotic cells.
Figures





Similar articles
-
Epigenomic modifications predict active promoters and gene structure in Toxoplasma gondii.PLoS Pathog. 2007 Jun;3(6):e77. doi: 10.1371/journal.ppat.0030077. PLoS Pathog. 2007. PMID: 17559302 Free PMC article.
-
Pair of unusual GCN5 histone acetyltransferases and ADA2 homologues in the protozoan parasite Toxoplasma gondii.Eukaryot Cell. 2006 Jan;5(1):62-76. doi: 10.1128/EC.5.1.62-76.2006. Eukaryot Cell. 2006. PMID: 16400169 Free PMC article.
-
Histone mediated gene activation in Toxoplasma gondii.Mol Biochem Parasitol. 2006 Aug;148(2):109-16. doi: 10.1016/j.molbiopara.2006.03.010. Epub 2006 Apr 18. Mol Biochem Parasitol. 2006. PMID: 16644030 Review.
-
A novel GCN5b lysine acetyltransferase complex associates with distinct transcription factors in the protozoan parasite Toxoplasma gondii.Mol Biochem Parasitol. 2019 Sep;232:111203. doi: 10.1016/j.molbiopara.2019.111203. Epub 2019 Aug 2. Mol Biochem Parasitol. 2019. PMID: 31381949 Free PMC article.
-
Toxoplasma histone acetylation remodelers as novel drug targets.Expert Rev Anti Infect Ther. 2012 Oct;10(10):1189-201. doi: 10.1586/eri.12.100. Expert Rev Anti Infect Ther. 2012. PMID: 23199404 Free PMC article. Review.
Cited by
-
Epigenetics as Driver of Adaptation and Diversification in Microbial Eukaryotes.Front Genet. 2021 Mar 16;12:642220. doi: 10.3389/fgene.2021.642220. eCollection 2021. Front Genet. 2021. PMID: 33796133 Free PMC article. No abstract available.
-
Yeast three-hybrid screen identifies TgBRADIN/GRA24 as a negative regulator of Toxoplasma gondii bradyzoite differentiation.PLoS One. 2015 Mar 19;10(3):e0120331. doi: 10.1371/journal.pone.0120331. eCollection 2015. PLoS One. 2015. PMID: 25789621 Free PMC article.
-
Protein arginine methyltransferases in protozoan parasites.Parasitology. 2022 Apr;149(4):427-435. doi: 10.1017/S0031182021002043. Epub 2021 Dec 6. Parasitology. 2022. PMID: 35331350 Free PMC article. Review.
-
Epigenetics in Plasmodium: what do we really know?Eukaryot Cell. 2010 Aug;9(8):1150-8. doi: 10.1128/EC.00093-10. Epub 2010 Jun 18. Eukaryot Cell. 2010. PMID: 20562224 Free PMC article. Review.
-
Quinoline derivative MC1626, a putative GCN5 histone acetyltransferase (HAT) inhibitor, exhibits HAT-independent activity against Toxoplasma gondii.Antimicrob Agents Chemother. 2007 Mar;51(3):1109-11. doi: 10.1128/AAC.01256-06. Epub 2006 Dec 18. Antimicrob Agents Chemother. 2007. PMID: 17178801 Free PMC article.
References
-
- Ahn, H. J., S. Kim, and H. W. Nam. 2003. Molecular cloning of the 82-kDa heat shock protein (HSP90) of Toxoplasma gondii associated with the entry into and growth in host cells. Biochem. Biophys. Res. Commun. 311:654-659. - PubMed
-
- An, W., J. Kim, and R. G. Roeder. 2004. Ordered cooperative functions of PRMT1, p300, and CARM1 in transcriptional activation by p53. Cell 117:735-748. - PubMed
-
- Bassi, M. T., R. S. Ramesar, B. Caciotti, I. M. Winship, A. De Grandi, M. Riboni, P. L. Townes, P. Beighton, A. Ballabio, and G. Borsani. 1999. X-linked late-onset sensorineural deafness caused by a deletion involving OA1 and a novel gene containing WD-40 repeats. Am. J. Hum. Genet. 64:1604-1616. - PMC - PubMed
-
- Bhatti, M. M., and W. J. Sullivan, Jr. 2005. Histone acetylase GCN5 enters the nucleus via importin-alpha in protozoan parasite Toxoplasma gondii. J. Biol. Chem. 280:5902-5908. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
