Sumoylation of p45/NF-E2: nuclear positioning and transcriptional activation of the mammalian beta-like globin gene locus
- PMID: 16287851
- PMCID: PMC1291221
- DOI: 10.1128/MCB.25.23.10365-10378.2005
Sumoylation of p45/NF-E2: nuclear positioning and transcriptional activation of the mammalian beta-like globin gene locus
Abstract
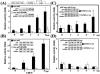
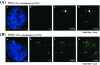
NF-E2 is a transcription activator for the regulation of a number of erythroid- and megakaryocytic lineage-specific genes. Here we present evidence that the large subunit of mammalian NF-E2, p45, is sumoylated in vivo in human erythroid K562 cells and in mouse fetal liver. By in vitro sumoylation reaction and DNA transfection experiments, we show that the sumoylation occurs at lysine 368 (K368) of human p45/NF-E2. Furthermore, p45 sumoylation enhances the transactivation capability of NF-E2, and this is accompanied by an increase of the NF-E2 DNA binding affinity. More interestingly, we have found that in K562 cells, the beta-globin gene loci in the euchromatin regions are predominantly colocalized with the nuclear bodies promyelocytic leukemia protein (PML) oncogenic domains that are enriched with the PML, SUMO-1, RNA polymerase II, and sumoylatable p45/NF-E2. Chromatin immunoprecipitation assays further showed that the intact sumoylation site of p45/NF-E2 is required for its binding to the DNase I-hypersensitive sites of the beta-globin locus control region. Finally, we demonstrated by stable transfection assay that only the wild-type p45, but not its mutant form p45 (K368R), could efficiently rescue beta-globin gene expression in the p45-null, erythroid cell line CB3. These data together point to a model of mammalian beta-like globin gene activation by sumoylated p45/NF-E2 in erythroid cells.
Figures








Similar articles
-
Chromatin-binding in vivo of the erythroid kruppel-like factor, EKLF, in the murine globin loci.Cell Res. 2006 Apr;16(4):347-55. doi: 10.1038/sj.cr.7310045. Cell Res. 2006. PMID: 16617330
-
Mouse microcytic anaemia caused by a defect in the gene encoding the globin enhancer-binding protein NF-E2.Nature. 1993 Apr 22;362(6422):768-70. doi: 10.1038/362768a0. Nature. 1993. PMID: 8469289
-
Heme-dependent up-regulation of the alpha-globin gene expression by transcriptional repressor Bach1 in erythroid cells.Biochem Biophys Res Commun. 2004 Nov 5;324(1):77-85. doi: 10.1016/j.bbrc.2004.09.022. Biochem Biophys Res Commun. 2004. PMID: 15464985
-
[Structure of genomic domains of mammalian and avian globin genes].Genetika. 2002 Dec;38(12):1589-606. Genetika. 2002. PMID: 12575443 Review. Russian.
-
The role of transcription factor NF-E2 in megakaryocyte maturation and platelet production.Stem Cells. 1996;14 Suppl 1:112-5. doi: 10.1002/stem.5530140714. Stem Cells. 1996. PMID: 11012210 Review.
Cited by
-
Sumoylation regulates interaction of FOG1 with C-terminal-binding protein (CTBP).J Biol Chem. 2010 Sep 3;285(36):28064-75. doi: 10.1074/jbc.M109.096909. Epub 2010 Jun 28. J Biol Chem. 2010. PMID: 20587419 Free PMC article.
-
Phosphorylation-dependent SUMOylation of the transcription factor NF-E2.PLoS One. 2012;7(9):e44608. doi: 10.1371/journal.pone.0044608. Epub 2012 Sep 10. PLoS One. 2012. PMID: 22970264 Free PMC article.
-
Prolonged fasting and glucocorticoid exposure drive dynamic DNA methylation in northern elephant seals.J Exp Biol. 2025 Aug 1;228(15):jeb250046. doi: 10.1242/jeb.250046. Epub 2025 Jul 25. J Exp Biol. 2025. PMID: 40583573 Free PMC article.
-
SUMOylation regulates the transcriptional repression activity of FOG-2 and its association with GATA-4.PLoS One. 2012;7(11):e50637. doi: 10.1371/journal.pone.0050637. Epub 2012 Nov 30. PLoS One. 2012. PMID: 23226341 Free PMC article.
-
Concomitant constitutive LNK and NFE2 mutation with loss of sumoylation in a case of hereditary thrombocythemia.Haematologica. 2021 Apr 1;106(4):1158-1162. doi: 10.3324/haematol.2020.246587. Haematologica. 2021. PMID: 32554556 Free PMC article. No abstract available.
References
-
- Amrolia, P. J., L. Ramamurthy, D. Saluja, N. Tanese, S. M. Jane, and J. M. Cunningham. 1997. The activation domain of the enhancer binding protein p45NF-E2 interacts with TAFII130 and mediates long-range activation of the alpha- and beta-globin gene loci in an erythroid cell line. Proc. Natl. Acad. Sci. USA 94:10051-10056. - PMC - PubMed
-
- Andrews, N. C. 1998. The NF-E2 transcription factor. Int. J. Biochem. Cell Biol. 30:429-432. - PubMed
-
- Andrews, N. C., H. Erdjument-Bromage, M. B. Davidson, P. Tempst, and S. H. Orkin. 1993. Erythroid transcription factor NF-E2 is a haematopoietic-specific basic-leucine zipper protein. Nature 362:722-728. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
