IMGT, the international ImMunoGeneTics information system: a standardized approach for immunogenetics and immunoinformatics
- PMID: 16305737
- PMCID: PMC1312312
- DOI: 10.1186/1745-7580-1-3
IMGT, the international ImMunoGeneTics information system: a standardized approach for immunogenetics and immunoinformatics
Abstract
IMGT, the international ImMunoGeneTics information system http://imgt.cines.fr, was created in 1989 by the Laboratoire d'ImmunoGénétique Moléculaire (LIGM) (Université Montpellier II and CNRS) at Montpellier, France. IMGT is a high quality integrated knowledge resource specialized in immunoglobulins (IG), T cell receptors (TR), major histocompatibility complex (MHC) of human and other vertebrates, and related proteins of the immune system (RPI) of any species which belong to the immunoglobulin superfamily (IgSF) and to the MHC superfamily (MhcSF). IMGT consists of five databases, ten on-line tools and more than 8,000 HTML pages of Web resources. IMGT provides a common access to standardized data from genome, genetics, proteome and three-dimensional structures. The accuracy and the consistency of IMGT data are based on IMGT-ONTOLOGY, a semantic specification of terms to be used in immunogenetics and immunoinformatics. IMGT-ONTOLOGY comprises six main concepts: IDENTIFICATION, CLASSIFICATION, DESCRIPTION, NUMEROTATION, ORIENTATION and OBTENTION. Based on these concepts, the controlled vocabulary and the annotation rules necessary for the immunogenetics data identification, classification, description and numbering and for the management of IMGT knowledge are defined in the IMGT Scientific chart. IMGT is the international reference in immunogenetics and immunoinformatics for medical research (repertoire analysis of the IG antibody sites and of the TR recognition sites in autoimmune and infectious diseases, AIDS, leukemias, lymphomas, myelomas), veterinary research (IG and TR repertoires in farm and wild life species), genome diversity and genome evolution studies of the adaptive immune responses, biotechnology related to antibody engineering (single chain Fragment variable (scFv), phage displays, combinatorial libraries, chimeric, humanized and human antibodies), diagnostics (detection and follow up of residual diseases) and therapeutical approaches (grafts, immunotherapy, vaccinology). IMGT is freely available at http://imgt.cines.fr.
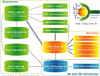
Figures


Similar articles
-
IMGT-Choreography for immunogenetics and immunoinformatics.In Silico Biol. 2005;5(1):45-60. In Silico Biol. 2005. PMID: 15972004
-
IMGT, the international ImMunoGeneTics information system.Nucleic Acids Res. 2005 Jan 1;33(Database issue):D593-7. doi: 10.1093/nar/gki065. Nucleic Acids Res. 2005. PMID: 15608269 Free PMC article.
-
IMGT-ONTOLOGY and IMGT databases, tools and Web resources for immunogenetics and immunoinformatics.Mol Immunol. 2004 Jan;40(10):647-60. doi: 10.1016/j.molimm.2003.09.006. Mol Immunol. 2004. PMID: 14644091 Review.
-
IMGT, the international ImMunoGeneTics information system, http://imgt.cines.fr: the reference in immunoinformatics.Stud Health Technol Inform. 2003;95:74-9. Stud Health Technol Inform. 2003. PMID: 14663966
-
IMGT, the international ImMunoGeneTics information system, http://imgt.cines.fr.Novartis Found Symp. 2003;254:126-36; discussion 136-42, 216-22, 250-2. Novartis Found Symp. 2003. PMID: 14712935 Review.
Cited by
-
IMGT®Homo sapiens IG and TR Loci, Gene Order, CNV and Haplotypes: New Concepts as a Paradigm for Jawed Vertebrates Genome Assemblies.Biomolecules. 2022 Feb 28;12(3):381. doi: 10.3390/biom12030381. Biomolecules. 2022. PMID: 35327572 Free PMC article.
-
Limited maintenance of vaccine-induced simian immunodeficiency virus-specific CD8 T-cell receptor clonotypes after virus challenge.J Virol. 2008 Aug;82(15):7357-68. doi: 10.1128/JVI.00607-08. Epub 2008 May 28. J Virol. 2008. PMID: 18508897 Free PMC article.
-
TCR-MHC docking orientation: natural selection, or thymic selection?Immunol Res. 2008;41(3):267-94. doi: 10.1007/s12026-008-8040-2. Immunol Res. 2008. PMID: 18726714 Review.
-
NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and -B locus protein of known sequence.PLoS One. 2007 Aug 29;2(8):e796. doi: 10.1371/journal.pone.0000796. PLoS One. 2007. PMID: 17726526 Free PMC article.
-
Structures of a key interaction protein from the Trypanosoma brucei editosome in complex with single domain antibodies.J Struct Biol. 2011 Apr;174(1):124-36. doi: 10.1016/j.jsb.2010.10.007. Epub 2010 Oct 20. J Struct Biol. 2011. PMID: 20969962 Free PMC article.
References
-
- Lefranc M-P, Clément O, Kaas Q, Duprat E, Chastellan P, Coelho I, Combres K, Ginestoux C, Giudicelli V, Chaume D, Lefranc G. IMGT-Choreography for immunogenetics and immunoinformatics. In Silico Biology. 2004;5:0006. Epub http://www.bioinfo.de/isb/2004/05/0006/, In Silico Biology, 2005, 5, 45-60. PMID: 15972004. - PubMed
-
- Lefranc M-P, Lefranc G. The Immunoglobulin FactsBook. Academic Press, London, UK; 2001. p. 458.http://imgt.cines.fr/textes/IMGTindex/factsbook.html
-
- Lefranc M-P, Lefranc G. The T cell receptor FactsBook. Academic Press, London, UK; 2001. p. 398.http://imgt.cines.fr/textes/IMGTindex/factsbook.html
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
