The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity
- PMID: 16306590
- PMCID: PMC1316023
- DOI: 10.1128/JVI.79.24.15189-15198.2005
The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity
Abstract
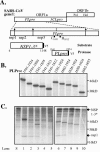
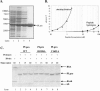
Replication of the genomic RNA of severe acute respiratory syndrome coronavirus (SARS-CoV) is mediated by replicase polyproteins that are processed by two viral proteases, papain-like protease (PLpro) and 3C-like protease (3CLpro). Previously, we showed that SARS-CoV PLpro processes the replicase polyprotein at three conserved cleavage sites. Here, we report the identification and characterization of a 316-amino-acid catalytic core domain of PLpro that can efficiently cleave replicase substrates in trans-cleavage assays and peptide substrates in fluorescent resonance energy transfer-based protease assays. We performed bioinformatics analysis on 16 papain-like protease domains from nine different coronaviruses and identified a putative catalytic triad (Cys1651-His1812-Asp1826) and zinc-binding site. Mutagenesis studies revealed that Asp1826 and the four cysteine residues involved in zinc binding are essential for SARS-CoV PLpro activity. Molecular modeling of SARS-CoV PLpro suggested that this catalytic core may also have deubiquitinating activity. We tested this hypothesis by measuring the deubiquitinating activity of PLpro by two independent assays. SARS CoV-PLpro hydrolyzed both diubiquitin and ubiquitin-7-amino-4-methylcoumarin (AMC) substrates, and hydrolysis of ubiquitin-AMC is approximately 180-fold more efficient than hydrolysis of a peptide substrate that mimics the PLpro replicase recognition sequence. To investigate the critical determinants recognized by PLpro, we performed site-directed mutagenesis on the P6 to P2' residues at each of the three PLpro cleavage sites. We found that PLpro recognizes the consensus cleavage sequence LXGG, which is also the consensus sequence recognized by cellular deubiquitinating enzymes. This similarity in the substrate recognition sites should be considered during the development of SARS-CoV PLpro inhibitors.
Figures



References
-
- Amerik, A. Y., and M. Hochstrasser. 2004. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta 1695:189-207. - PubMed
-
- Bacha, U., J. Barrila, A. Velazquez-Campoy, S. A. Leavitt, and E. Freire. 2004. Identification of novel inhibitors of the SARS coronavirus main protease 3CLpro. Biochemistry 43:4906-4912. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

