Contractile ring-independent localization of DdINCENP, a protein important for spindle stability and cytokinesis
- PMID: 16339076
- PMCID: PMC1356588
- DOI: 10.1091/mbc.e05-08-0704
Contractile ring-independent localization of DdINCENP, a protein important for spindle stability and cytokinesis
Abstract
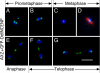
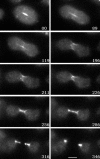
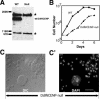
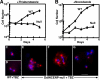
Dictyostelium DdINCENP is a chromosomal passenger protein associated with centromeres, the spindle midzone, and poles during mitosis and the cleavage furrow during cytokinesis. Disruption of the single DdINCENP gene revealed important roles for this protein in mitosis and cytokinesis. DdINCENP null cells lack a robust spindle midzone and are hypersensitive to microtubule-depolymerizing drugs, suggesting that their spindles may not be stable. Furthermore DdCP224, a protein homologous to the microtubule-stabilizing protein TOGp/XMAP215, was absent from the spindle midzone of DdINCENP null cells. Overexpression of DdCP224 rescued the weak spindle midzone defect of DdINCENP null cells. Although not required for the localization of the myosin II contractile ring and subsequent formation of a cleavage furrow, DdINCENP is important for the abscission of daughter cells at the end of cytokinesis. Finally, we show that the localization of DdINCENP at the cleavage furrow is modulated by myosin II but it occurs by a mechanism different from that controlling the formation of the contractile ring.
Figures





Similar articles
-
Dictyostelium Aurora kinase has properties of both Aurora A and Aurora B kinases.Eukaryot Cell. 2008 May;7(5):894-905. doi: 10.1128/EC.00422-07. Epub 2008 Mar 7. Eukaryot Cell. 2008. PMID: 18326585 Free PMC article.
-
The localization of inner centromeric protein (INCENP) at the cleavage furrow is dependent on Kif12 and involves interactions of the N terminus of INCENP with the actin cytoskeleton.Mol Biol Cell. 2007 Sep;18(9):3366-74. doi: 10.1091/mbc.e06-10-0895. Epub 2007 Jun 13. Mol Biol Cell. 2007. PMID: 17567958 Free PMC article.
-
A mitotic kinesin-like protein required for normal karyokinesis, myosin localization to the furrow, and cytokinesis in Dictyostelium.Proc Natl Acad Sci U S A. 2004 Nov 23;101(47):16519-24. doi: 10.1073/pnas.0407304101. Epub 2004 Nov 16. Proc Natl Acad Sci U S A. 2004. PMID: 15546981 Free PMC article.
-
Mechanics and regulation of cytokinesis.Curr Opin Cell Biol. 2004 Apr;16(2):182-8. doi: 10.1016/j.ceb.2004.02.002. Curr Opin Cell Biol. 2004. PMID: 15196562 Free PMC article. Review.
-
The role of centromere-binding factor 3 (CBF3) in spindle stability, cytokinesis, and kinetochore attachment.Biochem Cell Biol. 2005 Dec;83(6):696-702. doi: 10.1139/o05-161. Biochem Cell Biol. 2005. PMID: 16333320 Review.
Cited by
-
Dictyostelium centrin B localization during cell cycle progression.Commun Integr Biol. 2010 Jan;3(1):39-41. doi: 10.4161/cib.3.1.9721. Commun Integr Biol. 2010. PMID: 20539780 Free PMC article.
-
Genetic suppression of a phosphomimic myosin II identifies system-level factors that promote myosin II cleavage furrow accumulation.Mol Biol Cell. 2014 Dec 15;25(25):4150-65. doi: 10.1091/mbc.E14-08-1322. Epub 2014 Oct 15. Mol Biol Cell. 2014. PMID: 25318674 Free PMC article.
-
Dictyostelium Aurora kinase has properties of both Aurora A and Aurora B kinases.Eukaryot Cell. 2008 May;7(5):894-905. doi: 10.1128/EC.00422-07. Epub 2008 Mar 7. Eukaryot Cell. 2008. PMID: 18326585 Free PMC article.
-
Dynamic Mitotic Localization of the Centrosomal Kinases CDK1, Plk, AurK, and Nek2 in Dictyostelium amoebae.Cells. 2024 Sep 10;13(18):1513. doi: 10.3390/cells13181513. Cells. 2024. PMID: 39329697 Free PMC article.
-
The localization of inner centromeric protein (INCENP) at the cleavage furrow is dependent on Kif12 and involves interactions of the N terminus of INCENP with the actin cytoskeleton.Mol Biol Cell. 2007 Sep;18(9):3366-74. doi: 10.1091/mbc.e06-10-0895. Epub 2007 Jun 13. Mol Biol Cell. 2007. PMID: 17567958 Free PMC article.
References
-
- Adams, R. R., Carmena, M., and Earnshaw, W. C. (2001a). Chromosomal passengers and the (aurora) ABCs of mitosis. Trends Cell Biol. 11, 49-54. - PubMed
-
- Adams, R. R., Maiato, H., Earnshaw, W. C., and Carmena, M. (2001b). Essential roles of Drosophila inner centromere protein (INCENP) and aurora B in histone H3 phosphorylation, metaphase chromosome alignment, kinetochore disjunction, and chromosome segregation. J. Cell Biol. 153, 865-880. - PMC - PubMed
-
- Andrews, P. D., Ovechkina, Y., Morrice, N., Wagenbach, M., Duncan, K., Wordeman, L., and Swedlow, J. R. (2004). Aurora B regulates MCAK at the mitotic centromere. Dev. Cell 6, 253-268. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

