Formation of an energized inner membrane in mitochondria with a gamma-deficient F1-ATPase
- PMID: 16339725
- PMCID: PMC1317497
- DOI: 10.1128/EC.4.12.2078-2086.2005
Formation of an energized inner membrane in mitochondria with a gamma-deficient F1-ATPase
Abstract
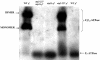
Eukaryotic cells require mitochondrial compartments for viability. However, the budding yeast Saccharomyces cerevisiae is able to survive when mitochondrial DNA suffers substantial deletions or is completely absent, so long as a sufficient mitochondrial inner membrane potential is generated. In the absence of functional mitochondrial DNA, and consequently a functional electron transport chain and F(1)F(o)-ATPase, the essential electrical potential is maintained by the electrogenic exchange of ATP(4-) for ADP(3-) through the adenine nucleotide translocator. An essential aspect of this electrogenic process is the conversion of ATP(4-) to ADP(3-) in the mitochondrial matrix, and the nuclear-encoded subunits of F(1)-ATPase are hypothesized to be required for this process in vivo. Deletion of ATP3, the structural gene for the gamma subunit of the F(1)-ATPase, causes yeast to quantitatively lose mitochondrial DNA and grow extremely slowly, presumably by interfering with the generation of an energized inner membrane. A spontaneous suppressor of this slow-growth phenotype was found to convert a conserved glycine to serine in the beta subunit of F(1)-ATPase (atp2-227). This mutation allowed substantial ATP hydrolysis by the F(1)-ATPase even in the absence of the gamma subunit, enabling yeast to generate a twofold greater inner membrane potential in response to ATP compared to mitochondria isolated from yeast lacking the gamma subunit and containing wild-type beta subunits. Analysis of the suppressing mutation by blue native polyacrylamide gel electrophoresis also revealed that the alpha(3)beta(3) heterohexamer can form in the absence of the gamma subunit.
Figures





Similar articles
-
F1-catalysed ATP hydrolysis is required for mitochondrial biogenesis in Saccharomyces cerevisiae growing under conditions where it cannot respire.Mol Microbiol. 2003 Mar;47(5):1329-39. doi: 10.1046/j.1365-2958.2003.03371.x. Mol Microbiol. 2003. PMID: 12603738
-
Genetic and biochemical basis for viability of yeast lacking mitochondrial genomes.Genetics. 2002 Dec;162(4):1595-604. doi: 10.1093/genetics/162.4.1595. Genetics. 2002. PMID: 12524335 Free PMC article.
-
Functional F1-ATPase essential in maintaining growth and membrane potential of human mitochondrial DNA-depleted rho degrees cells.J Biol Chem. 1998 Sep 4;273(36):22983-9. doi: 10.1074/jbc.273.36.22983. J Biol Chem. 1998. PMID: 9722521
-
The ADP and ATP transport in mitochondria and its carrier.Biochim Biophys Acta. 2008 Oct;1778(10):1978-2021. doi: 10.1016/j.bbamem.2008.04.011. Epub 2008 May 2. Biochim Biophys Acta. 2008. PMID: 18510943 Review.
-
Structure and function of the yeast vacuolar membrane proton ATPase.J Bioenerg Biomembr. 1989 Oct;21(5):589-603. doi: 10.1007/BF00808115. J Bioenerg Biomembr. 1989. PMID: 2531738 Review.
Cited by
-
ATP synthase: from single molecule to human bioenergetics.Proc Jpn Acad Ser B Phys Biol Sci. 2010;86(7):667-93. doi: 10.2183/pjab.86.667. Proc Jpn Acad Ser B Phys Biol Sci. 2010. PMID: 20689227 Free PMC article.
-
Ribosome-Targeting Antibiotics Impair T Cell Effector Function and Ameliorate Autoimmunity by Blocking Mitochondrial Protein Synthesis.Immunity. 2021 Jan 12;54(1):68-83.e6. doi: 10.1016/j.immuni.2020.11.001. Epub 2020 Nov 24. Immunity. 2021. PMID: 33238133 Free PMC article.
-
Phosphorylation of the F(1)F(o) ATP synthase beta subunit: functional and structural consequences assessed in a model system.Circ Res. 2010 Feb 19;106(3):504-13. doi: 10.1161/CIRCRESAHA.109.214155. Epub 2009 Dec 24. Circ Res. 2010. PMID: 20035080 Free PMC article.
-
Potential anti-aging agents suppress the level of constitutive mTOR- and DNA damage- signaling.Aging (Albany NY). 2012 Dec;4(12):952-65. doi: 10.18632/aging.100521. Aging (Albany NY). 2012. PMID: 23363784 Free PMC article.
-
Mutations in the Atp1p and Atp3p subunits of yeast ATP synthase differentially affect respiration and fermentation in Saccharomyces cerevisiae.J Bioenerg Biomembr. 2007 Apr;39(2):127-44. doi: 10.1007/s10863-007-9071-4. Epub 2007 May 10. J Bioenerg Biomembr. 2007. PMID: 17492370
References
-
- Abrahams, J. P., A. G. W. Leslie, R. Lutter, and J. Walker. 1994. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature (London) 370:621-628. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Appleby, R. D., W. K. Porteous, G. Hughes, A. M. James, D. Shannon, Y. H. Wei, and M. P. Murphy. 1999. Quantitation and origin of the mitochondrial membrane potential in human cells lacking mitochondrial DNA. Eur. J. Biochem. 262:108-116. - PubMed
-
- Baker, K. P., and G. Schatz. 1991. Mitochondrial proteins essential for viability mediate protein import into yeast mitochondria. Nature (London) 349:205-208. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

