Identification and characterization of Cor33p, a novel protein implicated in tolerance towards oxidative stress in Candida albicans
- PMID: 16339733
- PMCID: PMC1317491
- DOI: 10.1128/EC.4.12.2160-2169.2005
Identification and characterization of Cor33p, a novel protein implicated in tolerance towards oxidative stress in Candida albicans
Abstract
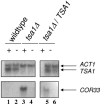
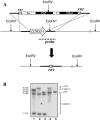
We applied two-dimensional gel electrophoresis to identify downstream effectors of CPH1 and EFG1 under hypha-inducing conditions in Candida albicans. Among the proteins that were expressed in wild-type cells but were strongly downregulated in a cph1Delta/efg1Delta double mutant in alpha-minimal essential medium at 37 degrees C, we could identify not-yet-characterized proteins, including Cor33-1p and Cor33-2p. The two proteins are almost identical (97% identity) and represent products of allelic isoforms of the same gene. Cor33p is highly similar to Cip1p from Candida sp. but lacks any significant homology to proteins from Saccharomyces cerevisiae. Strikingly, both proteins share homology with phenylcoumaran benzylic ether reductases and isoflavone reductases from plants. For other hypha-inducing media, like yeast-peptone-dextrose (YPD) plus serum at 37 degrees C, we could not detect any transcription for COR33 in wild-type cells, indicating that Cor33p is not hypha specific. In contrast, we found a strong induction for COR33 when cells were treated with 5 mM hydrogen peroxide. However, under oxidative conditions, transcription of COR33 was not dependent on EFG1, indicating that other regulatory factors are involved. In fact, upregulation depends on CAP1 at least, as transcript levels were clearly reduced in a Deltacap1 mutant strain under oxidative conditions. Unlike in wild-type cells, transcription of COR33 in a tsa1Delta mutant can be induced by treatment with 0.1 mM hydrogen peroxide. This suggests a functional link between COR33 and thiol-specific antioxidant-like proteins that are important in the oxidative-stress response in yeasts. Concordantly, cor33Delta deletion mutants show retarded growth on YPD plates supplemented with hydrogen peroxide, indicating that COR33 in general is implicated in conferring tolerance toward oxidative stress on Candida albicans.
Figures




Similar articles
-
Candida albicans response regulator gene SSK1 regulates a subset of genes whose functions are associated with cell wall biosynthesis and adaptation to oxidative stress.Eukaryot Cell. 2003 Oct;2(5):1018-24. doi: 10.1128/EC.2.5.1018-1024.2003. Eukaryot Cell. 2003. PMID: 14555484 Free PMC article.
-
Adaptive tolerance to oxidative stress and the induction of antioxidant enzymatic activities in Candida albicans are independent of the Hog1 and Cap1-mediated pathways.FEMS Yeast Res. 2010 Sep;10(6):747-56. doi: 10.1111/j.1567-1364.2010.00654.x. Epub 2010 Jun 7. FEMS Yeast Res. 2010. PMID: 20608985
-
Candida albicans Sfl1 suppresses flocculation and filamentation.Eukaryot Cell. 2007 Oct;6(10):1736-44. doi: 10.1128/EC.00236-07. Epub 2007 Aug 31. Eukaryot Cell. 2007. PMID: 17766464 Free PMC article.
-
Transcription factors regulating the response to oxidative stress in yeast.Antioxid Redox Signal. 2002 Feb;4(1):123-40. doi: 10.1089/152308602753625915. Antioxid Redox Signal. 2002. PMID: 11970849 Review.
-
Analysis of the Candida albicans proteome. II. Protein information technology on the Net (update 2002).J Chromatogr B Analyt Technol Biomed Life Sci. 2003 Apr 5;787(1):129-48. doi: 10.1016/s1570-0232(02)00762-6. J Chromatogr B Analyt Technol Biomed Life Sci. 2003. PMID: 12659738 Review.
Cited by
-
Normal adaptation of Candida albicans to the murine gastrointestinal tract requires Efg1p-dependent regulation of metabolic and host defense genes.Eukaryot Cell. 2013 Jan;12(1):37-49. doi: 10.1128/EC.00236-12. Epub 2012 Nov 2. Eukaryot Cell. 2013. PMID: 23125349 Free PMC article.
References
-
- Babiychuk, E., S. Kushnir, E. Belles-Boix, M. Van Montagu, and D. Inze. 1995. Arabidopsis thaliana NADPH oxidoreductase homologs confer tolerance of yeasts toward the thiol-oxidizing drug diamide. J. Biol. Chem. 270:26224-26231. - PubMed
-
- Braun, B. R., and A. D. Johnson. 1997. Control of filament formation in Candida albicans by the transcriptional repressor TUP1. Science 277:105-109. - PubMed
-
- Care, R. S., J. Trevethick, K. M. Binley, and P. E. Sudbery. 1999. The MET3 promoter: a new tool for Candida albicans molecular genetics. Mol. Microbiol. 34:792-798. - PubMed
-
- Dieterich, C., M. Schandar, M. Noll, F. J. Johannes, H. Brunner, T. Graeve, and S. Rupp. 2002. In vitro reconstructed human epithelia reveal contributions of Candida albicans EFG1 and CPH1 to adhesion and invasion. Microbiology 148:497-506. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

