Bacterial cell wall synthesis: new insights from localization studies
- PMID: 16339737
- PMCID: PMC1306805
- DOI: 10.1128/MMBR.69.4.585-607.2005
Bacterial cell wall synthesis: new insights from localization studies
Abstract
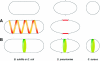
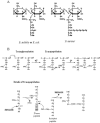
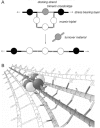
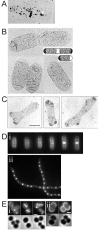
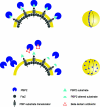
In order to maintain shape and withstand intracellular pressure, most bacteria are surrounded by a cell wall that consists mainly of the cross-linked polymer peptidoglycan (PG). The importance of PG for the maintenance of bacterial cell shape is underscored by the fact that, for various bacteria, several mutations affecting PG synthesis are associated with cell shape defects. In recent years, the application of fluorescence microscopy to the field of PG synthesis has led to an enormous increase in data on the relationship between cell wall synthesis and bacterial cell shape. First, a novel staining method enabled the visualization of PG precursor incorporation in live cells. Second, penicillin-binding proteins (PBPs), which mediate the final stages of PG synthesis, have been localized in various model organisms by means of immunofluorescence microscopy or green fluorescent protein fusions. In this review, we integrate the knowledge on the last stages of PG synthesis obtained in previous studies with the new data available on localization of PG synthesis and PBPs, in both rod-shaped and coccoid cells. We discuss a model in which, at least for a subset of PBPs, the presence of substrate is a major factor in determining PBP localization.
Figures

References
-
- Aarsman, M. E., A. Piette, C. Fraipont, T. M. Vinkenvleugel, M. Nguyen-Disteche, and T. den Blaauwen. 2005. Maturation of the Escherichia coli divisome occurs in two steps. Mol. Microbiol. 55:1631-1645. - PubMed
-
- Alaedini, A., and R. A. Day. 1999. Identification of two penicillin-binding multienzyme complexes in Haemophilus influenzae. Biochem. Biophys. Res. Commun. 264:191-195. - PubMed
-
- Asoh, S., H. Matsuzawa, F. Ishino, J. L. Strominger, M. Matsuhashi, and T. Ohta. 1986. Nucleotide sequence of the PBPA gene and characteristics of the deduced amino acid sequence of penicillin-binding protein 2 of Escherichia coli K12. Eur. J. Biochem. 160:231-238. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

