Assembly of CS1 pili: the role of specific residues of the major pilin, CooA
- PMID: 16352839
- PMCID: PMC1317577
- DOI: 10.1128/JB.188.1.231-239.2006
Assembly of CS1 pili: the role of specific residues of the major pilin, CooA
Abstract
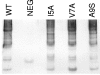
CS1 pili are important virulence factors of enterotoxigenic Escherichia coli strains associated with human diarrheal disease. They are the prototype for a family of pili that share extensive sequence similarity among their structural and assembly proteins. Only four linked genes, cooB, cooA, cooC, and cooD, are required to produce CS1 pili in E. coli K-12. To identify amino acids important for the function of the major pilin CooA, we used alanine substitution mutagenesis targeting conserved residues in the N and C termini of the protein. To test function, we examined cooA mutants for the ability to agglutinate bovine erythrocytes. Each hemagglutination-negative (HA(-)) cooA mutant was examined to identify its assembly pathway defect. CooA has been shown to be degraded in the absence of CooB (K. Voegele, H. Sakellaris, and J. R. Scott, Proc. Natl. Acad. Sci. USA 94:13257-13261, 1997). We found several HA(-) cooA mutants that produced no detectable CooA, suggesting that recognition by CooB is mediated by residues in both the N and C termini of CooA. In addition, we found that alanine substitution for some of the conserved residues in the C-terminal motif "AGxYxG(x(6))T," which is found in all subunits of this pilus family, had no effect on pilus formation. However, alanine substitution for some of the alternating hydrophobic residues within this motif prevented CooA from interacting with CooD, which serves as both the tip adhesin and nucleation protein for pilus formation. Thus, it appears that some, but not all, of the residues in both the N and C termini of CooA play a critical role in the intermolecular interactions of the major pilin with the other structural and assembly proteins. We anticipate that the results obtained here for CS1 pili in enterotoxigenic E. coli will help develop an understanding of the pilus assembly pathway used by CS1 family members in several important human pathogens.
Figures
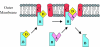

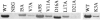



Similar articles
-
CooB plays a chaperone-like role for the proteins involved in formation of CS1 pili of enterotoxigenic Escherichia coli.Proc Natl Acad Sci U S A. 1997 Nov 25;94(24):13257-61. doi: 10.1073/pnas.94.24.13257. Proc Natl Acad Sci U S A. 1997. PMID: 9371833 Free PMC article.
-
A conserved residue in the tip proteins of CS1 and CFA/I pili of enterotoxigenic Escherichia coli that is essential for adherence.Proc Natl Acad Sci U S A. 1999 Oct 26;96(22):12828-32. doi: 10.1073/pnas.96.22.12828. Proc Natl Acad Sci U S A. 1999. PMID: 10536007 Free PMC article.
-
Assembly proteins of CS1 pili of enterotoxigenic Escherichia coli.Mol Microbiol. 1996 Aug;21(3):529-41. doi: 10.1111/j.1365-2958.1996.tb02562.x. Mol Microbiol. 1996. PMID: 8866477
-
New tools in an old trade: CS1 pilus morphogenesis.Mol Microbiol. 1998 Nov;30(4):681-7. doi: 10.1046/j.1365-2958.1998.01088.x. Mol Microbiol. 1998. PMID: 10094617 Review.
-
Chaperone-assisted assembly and molecular architecture of adhesive pili.Annu Rev Microbiol. 1991;45:383-415. doi: 10.1146/annurev.mi.45.100191.002123. Annu Rev Microbiol. 1991. PMID: 1683764 Review.
Cited by
-
Enterotoxigenic Escherichia coli CS1 pilus: not one structure but several.J Bacteriol. 2013 Apr;195(7):1357-9. doi: 10.1128/JB.00053-13. Epub 2013 Jan 25. J Bacteriol. 2013. PMID: 23354749 Free PMC article. No abstract available.
-
Discovery of bioactive microbial gene products in inflammatory bowel disease.Nature. 2022 Jun;606(7915):754-760. doi: 10.1038/s41586-022-04648-7. Epub 2022 May 25. Nature. 2022. PMID: 35614211 Free PMC article.
-
Adhesins Involved in Attachment to Abiotic Surfaces by Gram-Negative Bacteria.Microbiol Spectr. 2015 Aug;3(4):10.1128/microbiolspec.MB-0018-2015. doi: 10.1128/microbiolspec.MB-0018-2015. Microbiol Spectr. 2015. PMID: 26350310 Free PMC article. Review.
-
Transcriptional regulation of subclass 5b fimbriae.BMC Microbiol. 2008 Oct 14;8:180. doi: 10.1186/1471-2180-8-180. BMC Microbiol. 2008. PMID: 18854044 Free PMC article.
-
Functional Role of N- and C-Terminal Amino Acids in the Structural Subunits of Colonization Factor CS6 Expressed by Enterotoxigenic Escherichia coli.J Bacteriol. 2016 Apr 14;198(9):1429-41. doi: 10.1128/JB.00657-15. Print 2016 May. J Bacteriol. 2016. PMID: 26929298 Free PMC article.
References
-
- Amann, E., B. Ochs, and K. J. Abel. 1988. Tightly regulated tac promoter vectors useful for the expression of unfused and fused proteins in Escherichia coli. Gene 69:301-315. - PubMed
-
- Barry, E. M., Z. Altboum, G. Losonsky, and M. M. Levine. 2003. Immune responses elicited against multiple enterotoxigenic Escherichia coli fimbriae and mutant LT expressed in attenuated Shigella vaccine strains. Vaccine 21:333-340. - PubMed
-
- Black, R. E. 1993. Epidemiology of diarrhoeal disease: implications for control by vaccines. Vaccine 11:100-106. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

