The autoregulatory translational control element of poly(A)-binding protein mRNA forms a heteromeric ribonucleoprotein complex
- PMID: 16356927
- PMCID: PMC1316114
- DOI: 10.1093/nar/gki1014
The autoregulatory translational control element of poly(A)-binding protein mRNA forms a heteromeric ribonucleoprotein complex
Abstract
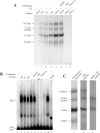
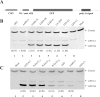
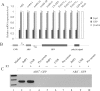
Repression of poly(A)-binding protein (PABP) mRNA translation involves the binding of PABP to the adenine-rich autoregulatory sequence (ARS) in the 5'-untranslated region of its own mRNA. In this report, we show that the ARS forms a complex in vitro with PABP, and two additional polypeptides of 63 and 105 kDa. The 63 and 105 kDa polypeptides were identified, as IMP1, an ortholog of chicken zip-code binding polypeptide, and UNR, a PABP binding polypeptide, respectively, by mass spectrometry of the ARS RNA affinity purified samples. Using a modified ribonucleoprotein (RNP) immunoprecipitation procedure we further show that indeed, both IMP1 and UNR bind to the ARS containing reporter RNA in vivo. Although both IMP1 and UNR could bind independently to the ARS RNA in vitro, their RNA-binding ability was stimulated by PABP. Mutational analyses of the ARS show that the presence of four of the six oligo(A) regions of the ARS was sufficient to repress translation and the length of the conserved pyrimidine spacers between the oligo(A) sequences was important for ARS function. The ability of mutant ARS RNAs to form the PABP, IMP1 and UNR containing RNP complex correlates well with the translational repressor activity of the ARS. There is also a direct relationship between the length of the poly(A) RNAs and their ability to form a trimeric complex with PABP, and to repress mRNA translation. UV crosslinking studies suggest that the ARS is less efficient than a poly(A) RNA of similar length, to bind to PABP. We show here that the ARS cannot efficiently form a trimeric complex with PABP; therefore, additional interactions with IMP1 and UNR to form a heteromeric RNP complex may be required for maximal repression of PABP mRNA translation under physiological conditions.
Figures




Similar articles
-
IMP1 interacts with poly(A)-binding protein (PABP) and the autoregulatory translational control element of PABP-mRNA through the KH III-IV domain.FEBS J. 2006 Dec;273(24):5678-90. doi: 10.1111/j.1742-4658.2006.05556.x. FEBS J. 2006. PMID: 17212783
-
Negative control of the poly(A)-binding protein mRNA translation is mediated by the adenine-rich region of its 5'-untranslated region.J Biol Chem. 1998 Dec 18;273(51):34535-42. doi: 10.1074/jbc.273.51.34535. J Biol Chem. 1998. PMID: 9852123
-
The SXL-UNR corepressor complex uses a PABP-mediated mechanism to inhibit ribosome recruitment to msl-2 mRNA.Mol Cell. 2009 Nov 25;36(4):571-82. doi: 10.1016/j.molcel.2009.09.042. Mol Cell. 2009. PMID: 19941818
-
Regulation of poly(A)-binding protein through PABP-interacting proteins.Cold Spring Harb Symp Quant Biol. 2006;71:537-43. doi: 10.1101/sqb.2006.71.061. Cold Spring Harb Symp Quant Biol. 2006. PMID: 17381337 Review.
-
LARP1 and LARP4: up close with PABP for mRNA 3' poly(A) protection and stabilization.RNA Biol. 2021 Feb;18(2):259-274. doi: 10.1080/15476286.2020.1868753. Epub 2021 Jan 31. RNA Biol. 2021. PMID: 33522422 Free PMC article. Review.
Cited by
-
A role for the poly(A)-binding protein Pab1p in PUF protein-mediated repression.J Biol Chem. 2011 Sep 23;286(38):33268-78. doi: 10.1074/jbc.M111.264572. Epub 2011 Jul 15. J Biol Chem. 2011. PMID: 21768112 Free PMC article.
-
Nuclear localization of cytoplasmic poly(A)-binding protein upon rotavirus infection involves the interaction of NSP3 with eIF4G and RoXaN.J Virol. 2008 Nov;82(22):11283-93. doi: 10.1128/JVI.00872-08. Epub 2008 Sep 17. J Virol. 2008. PMID: 18799579 Free PMC article.
-
A new role for the cellular PABP repressor Paip2 as an innate restriction factor capable of limiting productive cytomegalovirus replication.Genes Dev. 2013 Aug 15;27(16):1809-20. doi: 10.1101/gad.221341.113. Genes Dev. 2013. PMID: 23964095 Free PMC article.
-
CSDE1: a versatile regulator of gene expression in cancer.NAR Cancer. 2024 Apr 10;6(2):zcae014. doi: 10.1093/narcan/zcae014. eCollection 2024 Jun. NAR Cancer. 2024. PMID: 38600987 Free PMC article.
-
Sex determination in Drosophila: The view from the top.Fly (Austin). 2010 Jan-Mar;4(1):60-70. doi: 10.4161/fly.4.1.11277. Epub 2010 Jan 21. Fly (Austin). 2010. PMID: 20160499 Free PMC article. Review.
References
-
- Le H., Tanguay R.L., Balasta M.L., Wei C.C., Browning K.S., Metz A.M., Goss D.J., Gallie D.R. Translation initiation factors eIF-iso4G and eIF-4B interact with the poly(A)-binding protein and increase its RNA binding activity. J. Biol. Chem. 1997;272:16247–16255. - PubMed
-
- Cosson B., Berkova N., Couturier A., Chabelskaya S., Philippe M., Zhouravleva G. Poly(A)-binding protein and eRF3 are associated in vivo in human and Xenopus cells. Biol. Cell. 2002;94:205–216. - PubMed
-
- Craig A.W., Harhighat A., Yu A.T., Sonenberg N. Interaction of polyadenylate-binding protein with eIF4G homologue PAIP enhances translation. Nature. 1998;392:520–523. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

