Dynamic regulation of replication independent deposition of histone H3 in fission yeast
- PMID: 16361268
- PMCID: PMC1316113
- DOI: 10.1093/nar/gki1011
Dynamic regulation of replication independent deposition of histone H3 in fission yeast
Abstract
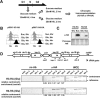
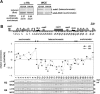
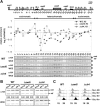
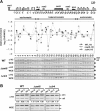
Recently, a histone H3 variant in Drosophila and humans, the H3.3 protein, was shown to replace canonical H3 in active chromatin in a replication-independent (RI) manner. In the fission yeast Schizosaccharomyces pombe, there exists a single form of H3, which is equivalent to H3.3 and is thought to participate in both replication-independent (RI) and replication-coupled (RC) nucleosome assembly. In this study, we show that RI deposition of H3 at heterochromatic regions is consistently lower than that at a gene-free euchromatic region, and deletion of the conserved heterochromatin-specific proteins Swi6 or Clr4 markedly increases RI deposition at heterochromatic regions such as the silent mating-type loci or centromeres. These results clearly show that RI deposition of H3 occurs preferentially in euchromatic regions. We also observed that RI deposition of H3 could be increased at the thi3(+) gene when transcription is induced, indicating transcription further facilitates RI deposition of H3. Taken together, these observations demonstrate that selective deposition of histone H3.3 at transcriptionally active chromatin by the RI assembly pathway is conserved in fission yeast and, thus, our data support an essential role of histone H3 replacement in maintaining active chromatin among diverse eukaryotic organisms ranging from fission yeast to humans.
Figures

References
-
- Workman J.L., Kingston R.E. Alteration of nucleosome structure as a mechanism of transcriptional regulation. Annu. Rev. Biochem. 1998;67:545–579. - PubMed
-
- Mello J.A., Almouzni G. The ins and outs of nucleosome assembly. Curr. Opin. Genet. Dev. 2001;11:136–141. - PubMed
-
- Ray-Gallet D., Quivy J.P., Scamps C., Martini E.M., Lipinski M., Almouzni G. HIRA is critical for a nucleosome assembly pathway independent of DNA synthesis. Mol. Cell. 2002;9:1091–1100. - PubMed
-
- Ahmad K., Henikoff S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell. 2002;9:1191–1200. - PubMed

