Single-molecule unfolding force distributions reveal a funnel-shaped energy landscape
- PMID: 16361331
- PMCID: PMC1367298
- DOI: 10.1529/biophysj.105.077982
Single-molecule unfolding force distributions reveal a funnel-shaped energy landscape
Abstract
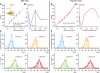
The protein folding process is described as diffusion on a high-dimensional energy landscape. Experimental data showing details of the underlying energy surface are essential to understanding folding. So far in single-molecule mechanical unfolding experiments a simplified model assuming a force-independent transition state has been used to extract such information. Here we show that this so-called Bell model, although fitting well to force velocity data, fails to reproduce full unfolding force distributions. We show that by applying Kramers' diffusion model, we were able to reconstruct a detailed funnel-like curvature of the underlying energy landscape and establish full agreement with the data. We demonstrate that obtaining spatially resolved details of the unfolding energy landscape from mechanical single-molecule protein unfolding experiments requires models that go beyond the Bell model.
Figures
Similar articles
-
Optimizing the calculation of energy landscape parameters from single-molecule protein unfolding experiments.Phys Rev E Stat Nonlin Soft Matter Phys. 2015 Jan;91(1):012710. doi: 10.1103/PhysRevE.91.012710. Epub 2015 Jan 26. Phys Rev E Stat Nonlin Soft Matter Phys. 2015. PMID: 25679645
-
Mapping the energy landscape of biomolecules using single molecule force correlation spectroscopy: theory and applications.Biophys J. 2006 Jun 1;90(11):3827-41. doi: 10.1529/biophysj.105.075937. Epub 2006 Mar 13. Biophys J. 2006. PMID: 16533852 Free PMC article.
-
Role of pulling direction in understanding the energy landscape of proteins.Phys Rev E Stat Nonlin Soft Matter Phys. 2008 Aug;78(2 Pt 1):021905. doi: 10.1103/PhysRevE.78.021905. Epub 2008 Aug 13. Phys Rev E Stat Nonlin Soft Matter Phys. 2008. PMID: 18850863
-
From folding theories to folding proteins: a review and assessment of simulation studies of protein folding and unfolding.Annu Rev Phys Chem. 2001;52:499-535. doi: 10.1146/annurev.physchem.52.1.499. Annu Rev Phys Chem. 2001. PMID: 11326073 Review.
-
The experimental survey of protein-folding energy landscapes.Q Rev Biophys. 2005 Aug;38(3):245-88. doi: 10.1017/S0033583506004185. Epub 2006 Jun 19. Q Rev Biophys. 2005. PMID: 16780604 Review.
Cited by
-
Hidden multiple bond effects in dynamic force spectroscopy.Biophys J. 2012 Mar 7;102(5):1184-93. doi: 10.1016/j.bpj.2012.01.037. Epub 2012 Mar 6. Biophys J. 2012. PMID: 22404941 Free PMC article.
-
Complex unfolding kinetics of single-domain proteins in the presence of force.Biophys J. 2010 Sep 8;99(5):1620-7. doi: 10.1016/j.bpj.2010.06.039. Biophys J. 2010. PMID: 20816075 Free PMC article.
-
Affinity-matured recombinant antibody fragments analyzed by single-molecule force spectroscopy.Biophys J. 2007 Nov 15;93(10):3583-90. doi: 10.1529/biophysj.107.112532. Epub 2007 Aug 3. Biophys J. 2007. PMID: 17675348 Free PMC article.
-
The effects of macromolecular crowding on the mechanical stability of protein molecules.Protein Sci. 2008 Dec;17(12):2156-66. doi: 10.1110/ps.037325.108. Epub 2008 Sep 9. Protein Sci. 2008. PMID: 18780817 Free PMC article.
-
Effects of multiple-bond ruptures on kinetic parameters extracted from force spectroscopy measurements: revisiting biotin-streptavidin interactions.Biophys J. 2008 Oct;95(8):3964-76. doi: 10.1529/biophysj.108.133900. Epub 2008 Jul 11. Biophys J. 2008. PMID: 18621812 Free PMC article.
References
-
- Onuchic, J. N., and P. G. Wolynes. 2004. Theory of protein folding. Curr. Opin. Struct. Biol. 14:70–75. - PubMed
-
- Carrion-Vazquez, M., A. F. Oberhauser, T. E. Fisher, P. E. Marszalek, H. Li, and J. M. Fernandez. 2000. Mechanical design of proteins studied by single-molecule force spectroscopy and protein engineering. Prog. Biophys. Mol. Biol. 74:63–91. - PubMed
-
- Merkel, R., P. Nassoy, A. Leung, K. Ritchie, and E. Evans. 1999. Energy landscapes of receptor-ligand bonds explored with dynamic force spectroscopy. Nature. 397:50–53. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

