Lectin receptor kinases participate in protein-protein interactions to mediate plasma membrane-cell wall adhesions in Arabidopsis
- PMID: 16361528
- PMCID: PMC1326033
- DOI: 10.1104/pp.105.066464
Lectin receptor kinases participate in protein-protein interactions to mediate plasma membrane-cell wall adhesions in Arabidopsis
Abstract
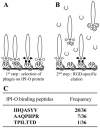
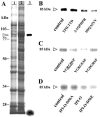
Interactions between plant cell walls and plasma membranes are essential for cells to function properly, but the molecules that mediate the structural continuity between wall and membrane are unknown. Some of these interactions, which are visualized upon tissue plasmolysis in Arabidopsis (Arabidopsis thaliana), are disrupted by the RGD (arginine-glycine-aspartic acid) tripeptide sequence, a characteristic cell adhesion motif in mammals. In planta induced-O (IPI-O) is an RGD-containing protein from the plant pathogen Phytophthora infestans that can disrupt cell wall-plasma membrane adhesions through its RGD motif. To identify peptide sequences that specifically bind the RGD motif of the IPI-O protein and potentially play a role in receptor recognition, we screened a heptamer peptide library displayed in a filamentous phage and selected two peptides acting as inhibitors of the plasma membrane RGD-binding activity of Arabidopsis. Moreover, the two peptides also disrupted cell wall-plasma membrane adhesions. Sequence comparison of the RGD-binding peptides with the Arabidopsis proteome revealed 12 proteins containing amino acid sequences in their extracellular domains common with the two RGD-binding peptides. Eight belong to the receptor-like kinase family, four of which have a lectin-like extracellular domain. The lectin domain of one of these, At5g60300, recognized the RGD motif both in peptides and proteins. These results imply that lectin receptor kinases are involved in protein-protein interactions with RGD-containing proteins as potential ligands, and play a structural and signaling role at the plant cell surfaces.
Figures

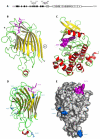

References
-
- Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796–815 - PubMed
-
- Bardy N, Carrasco A, Galaud JP, Pont-Lezica R, Canut H (1998) Free-flow electrophoresis for fractionation of Arabidopsis membranes. Electrophoresis 19: 1145–1153 - PubMed
-
- Barre A, Hervé C, Lescure B, Rougé P (2002) Lectin receptor kinases in plants. CRC Crit Rev Plant Sci 21: 379–399
-
- Berger F, Taylor A, Brownlee C (1994) Cell fate determination by the cell wall in early Fucus development. Science 263: 1421–1423 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

