Protein stabilization by specific binding of guanidinium to a functional arginine-binding surface on an SH3 domain
- PMID: 16373478
- PMCID: PMC2242360
- DOI: 10.1110/ps.051829106
Protein stabilization by specific binding of guanidinium to a functional arginine-binding surface on an SH3 domain
Abstract
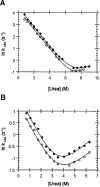
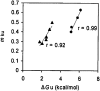
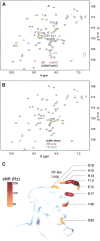
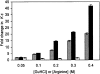
Guanidinium hydrochloride (GuHCl) at low concentrations significantly stabilizes the Fyn SH3 domain. In this work, we have demonstrated that this stabilizing effect is manifested through a dramatic (five- to sixfold) decrease in the unfolding rate of the domain with the folding rate being affected minimally. This behavior contrasts to the effect of NaCl, which stabilizes this domain by accelerating the folding rate. These data imply that the stabilizing effect of GuHCl is not predominantly ionic in nature. Through NMR studies, we have identified a specific binding site for guanidinium, and we have determined a dissociation constant of 90 mM for this interaction. The guanidinium-binding site overlaps with a functionally important arginine-binding pocket on the domain surface, and we have shown that GuHCl is a specific inhibitor of the peptide-binding activity of the domain. A different SH3 domain possessing a similar arginine-binding pocket is also thermodynamically stabilized by GuHCl. These data suggest that many proteins that normally interact with arginine-containing ligands may also be able to specifically interact with guanidinium. Thus, some caution should be used when using GuHCl as a denaturant in protein folding studies. Since arginine-mediated interactions are often important in the energetics of protein-protein interactions, our observations could be relevant for the design of small molecule inhibitors of protein-protein interactions.
Figures



Similar articles
-
Apparent Debye-Huckel electrostatic effects in the folding of a simple, single domain protein.Biochemistry. 2005 Feb 1;44(4):1243-50. doi: 10.1021/bi048444l. Biochemistry. 2005. PMID: 15667218
-
The role of backbone motions in ligand binding to the c-Src SH3 domain.J Mol Biol. 2001 Nov 2;313(4):873-87. doi: 10.1006/jmbi.2001.5083. J Mol Biol. 2001. PMID: 11697910
-
Interactions between the Fyn SH3-domain and adaptor protein Cbp/PAG derived ligands, effects on kinase activity and affinity.FEBS J. 2008 Oct;275(19):4863-74. doi: 10.1111/j.1742-4658.2008.06626.x. Epub 2008 Aug 21. FEBS J. 2008. PMID: 18721137
-
Probing invisible, low-populated States of protein molecules by relaxation dispersion NMR spectroscopy: an application to protein folding.Acc Chem Res. 2008 Mar;41(3):442-51. doi: 10.1021/ar700189y. Epub 2008 Feb 15. Acc Chem Res. 2008. PMID: 18275162 Review.
-
Is arginine a protein-denaturant?Protein Expr Purif. 2005 Jul;42(1):1-6. doi: 10.1016/j.pep.2005.03.028. Epub 2005 Apr 19. Protein Expr Purif. 2005. PMID: 15893471 Review.
Cited by
-
Stabilization Effect of Intrinsically Disordered Regions on Multidomain Proteins: The Case of the Methyl-CpG Protein 2, MeCP2.Biomolecules. 2021 Aug 16;11(8):1216. doi: 10.3390/biom11081216. Biomolecules. 2021. PMID: 34439881 Free PMC article.
-
NMR Dynamic View of the Stabilization of the WW4 Domain by Neutral NaCl and Kosmotropic Na2SO4 and NaH2PO4.Int J Mol Sci. 2024 Aug 22;25(16):9091. doi: 10.3390/ijms25169091. Int J Mol Sci. 2024. PMID: 39201778 Free PMC article.
-
The Recombinant Inhibitor of DNA Binding Id2 Forms Multimeric Structures via the Helix-Loop-Helix Domain and the Nuclear Export Signal.Int J Mol Sci. 2018 Apr 7;19(4):1105. doi: 10.3390/ijms19041105. Int J Mol Sci. 2018. PMID: 29642431 Free PMC article.
-
Does Microsecond Active-Site Dynamics Primarily control Proteolytic Activity of Bromelain? Clues from Single Molecular Level Study with a Denaturant, a Stabilizer and a Macromolecular Crowder.BBA Adv. 2022 Jan 12;2:100041. doi: 10.1016/j.bbadva.2022.100041. eCollection 2022. BBA Adv. 2022. PMID: 37082607 Free PMC article.
-
Disassembly of Single Virus Capsids Monitored in Real Time with Multicycle Resistive-Pulse Sensing.Anal Chem. 2022 Jan 18;94(2):985-992. doi: 10.1021/acs.analchem.1c03855. Epub 2021 Dec 21. Anal Chem. 2022. PMID: 34932317 Free PMC article.
References
-
- Bhuyan, A.K. 2002. Protein stabilization by urea and guanidine hydrochloride. Biochemistry 41: 13386–13394. - PubMed
-
- Bogan, A.A. and Thorn, K.S. 1998. Anatomy of hot spots in protein interfaces. J. Mol. Biol. 280: 1–9. - PubMed
-
- Collins, K.D. 2004. Ions from the Hofmeister series and osmolytes: Effects on proteins in solution and in the crystallization process. Methods 34: 300–311. - PubMed
-
- de Los Rios, M.A. and Plaxco, K.W. 2005. Apparent Debye-Huckel electrostatic effects in the folding of a simple, single domain protein. Biochemistry 44: 1243–1250. - PubMed
-
- Delaglio, F., Grzesiek, S., Vuister, G.W., Zhu, G., Pfeifer, J., and Bax, A. 1995. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6: 277–293. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

