The E7 proteins of low- and high-risk human papillomaviruses share the ability to target the pRB family member p130 for degradation
- PMID: 16381817
- PMCID: PMC1326189
- DOI: 10.1073/pnas.0510012103
The E7 proteins of low- and high-risk human papillomaviruses share the ability to target the pRB family member p130 for degradation
Abstract
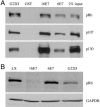
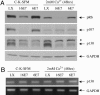
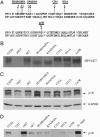
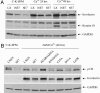
High-risk human papillomaviruses (HPVs) (e.g., HPV-16) cause anogenital and head and neck cancers, and low-risk HPVs (e.g., HPV-6) cause benign hyperproliferative disease. The E7 protein of HPV-16 binds all retinoblastoma tumor suppressor protein (pRB) family members with higher affinity than HPV-6E7. The HPV-16 E7 protein has been reported to target pRB family members for degradation and to immortalize cells. Here we tested the hypothesis that the low-risk E7 protein has an intrinsic ability to decrease expression of pRB family members. First, we introduced a high-affinity pRB-binding site into HPV-6 E7 (6E7G22D) and showed that, in human foreskin keratinocytes, HPV-6 E7G22D decreased the level of pRB protein but not pRB mRNA. Second, we analyzed the ability of wild-type HPV-6 E7 to destabilize the other pRB family members, p107 and p130. HPV-6 E7, like HPV-16 E7, decreased the level of p130 protein. This decrease was inhibited by MG132, a proteasome inhibitor. Binding of HPV-6 E7 to p130 was necessary but not sufficient to decrease the level of p130. Furthermore, the destabilization of p130 correlated with a decrease in the expression of involucrin, a differentiation marker. We suggest that the shared activity of HPV-16 E7 and HPV-6 E7 to destabilize p130 and decrease or delay differentiation may be related to the role of E7 in the HPV life cycle. The added ability of HPV-16 E7 to regulate pRB and p107 may be related to oncogenic activity.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

