MODOMICS: a database of RNA modification pathways
- PMID: 16381833
- PMCID: PMC1347447
- DOI: 10.1093/nar/gkj084
MODOMICS: a database of RNA modification pathways
Abstract
MODOMICS is the first comprehensive database resource for systems biology of RNA modification. It integrates information about the chemical structure of modified nucleosides, their localization in RNA sequences, pathways of their biosynthesis and enzymes that carry out the respective reactions. MODOMICS also provides literature information, and links to other databases, including the available protein sequence and structure data. The current list of modifications and pathways is comprehensive, while the dataset of enzymes is limited to Escherichia coli and Saccharomyces cerevisiae and sequence alignments are presented only for tRNAs from these organisms. RNAs and enzymes from other organisms will be included in the near future. MODOMICS can be queried by the type of nucleoside (e.g. A, G, C, U, I, m1A, nm5s2U, etc.), type of RNA, position of a particular nucleoside, type of reaction (e.g. methylation, thiolation, deamination, etc.) and name or sequence of an enzyme of interest. Options for data presentation include graphs of pathways involving the query nucleoside, multiple sequence alignments of RNA sequences and tabular forms with enzyme and literature data. The contents of MODOMICS can be accessed through the World Wide Web at http://genesilico.pl/modomics/.
Figures
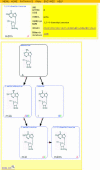

References
-
- Grosjean H., Benne R. Modification and Editing of RNA. Washington, DC: ASM Press; 1998.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

