The YEASTRACT database: a tool for the analysis of transcription regulatory associations in Saccharomyces cerevisiae
- PMID: 16381908
- PMCID: PMC1347376
- DOI: 10.1093/nar/gkj013
The YEASTRACT database: a tool for the analysis of transcription regulatory associations in Saccharomyces cerevisiae
Abstract
We present the YEAst Search for Transcriptional Regulators And Consensus Tracking (YEASTRACT; www.yeastract.com) database, a tool for the analysis of transcription regulatory associations in Saccharomyces cerevisiae. This database is a repository of 12 346 regulatory associations between transcription factors and target genes, based on experimental evidence which was spread throughout 861 bibliographic references. It also includes 257 specific DNA-binding sites for more than a hundred characterized transcription factors. Further information about each yeast gene included in the database was obtained from Saccharomyces Genome Database (SGD), Regulatory Sequences Analysis Tools and Gene Ontology (GO) Consortium. Computational tools are also provided to facilitate the exploitation of the gathered data when solving a number of biological questions as exemplified in the Tutorial also available on the system. YEASTRACT allows the identification of documented or potential transcription regulators of a given gene and of documented or potential regulons for each transcription factor. It also renders possible the comparison between DNA motifs, such as those found to be over-represented in the promoter regions of co-regulated genes, and the transcription factor-binding sites described in the literature. The system also provides an useful mechanism for grouping a list of genes (for instance a set of genes with similar expression profiles as revealed by microarray analysis) based on their regulatory associations with known transcription factors.
Figures
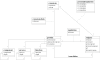
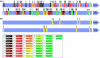
References
-
- Goffeau A., Barrell B., Bussey H., Davis R.W., Dujon B., Feldmann H., Galibert F., Hoheisel J., Jacq C., Johnston M., et al. Life with 6000 genes. Science. 1996;274:563–567. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

