Characterization of PhlG, a hydrolase that specifically degrades the antifungal compound 2,4-diacetylphloroglucinol in the biocontrol agent Pseudomonas fluorescens CHA0
- PMID: 16391073
- PMCID: PMC1352262
- DOI: 10.1128/AEM.72.1.418-427.2006
Characterization of PhlG, a hydrolase that specifically degrades the antifungal compound 2,4-diacetylphloroglucinol in the biocontrol agent Pseudomonas fluorescens CHA0
Abstract
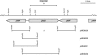
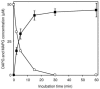
The potent antimicrobial compound 2,4-diacetylphloroglucinol (DAPG) is a major determinant of biocontrol activity of plant-beneficial Pseudomonas fluorescens CHA0 against root diseases caused by fungal pathogens. The DAPG biosynthetic locus harbors the phlG gene, the function of which has not been elucidated thus far. The phlG gene is located upstream of the phlACBD biosynthetic operon, between the phlF and phlH genes which encode pathway-specific regulators. In this study, we assigned a function to PhlG as a hydrolase specifically degrades DAPG to equimolar amounts of mildly toxic monoacetylphloroglucinol (MAPG) and acetate. DAPG added to cultures of a DAPG-negative DeltaphlA mutant of strain CHA0 was completely degraded, and MAPG was temporarily accumulated. In contrast, DAPG was not degraded in cultures of a DeltaphlA DeltaphlG double mutant. To confirm the enzymatic nature of PhlG in vitro, the protein was histidine tagged, overexpressed in Escherichia coli, and purified by affinity chromatography. Purified PhlG had a molecular mass of about 40 kDa and catalyzed the degradation of DAPG to MAPG. The enzyme had a kcat of 33 s(-1) and a Km of 140 microM at 30 degrees C and pH 7. The PhlG enzyme did not degrade other compounds with structures similar to DAPG, such as MAPG and triacetylphloroglucinol, suggesting strict substrate specificity. Interestingly, PhlG activity was strongly reduced by pyoluteorin, a further antifungal compound produced by the bacterium. Expression of phlG was not influenced by the substrate DAPG or the degradation product MAPG but was subject to positive control by the GacS/GacA two-component system and to negative control by the pathway-specific regulators PhlF and PhlH.
Figures
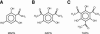
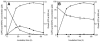

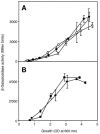
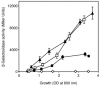
References
-
- Abbas, A., J. E. McGuire, D. Crowley, C. Baysse, M. Dow, and F. O'Gara. 2004. The putative permease PhlE of Pseudomonas fluorescens F113 has a role in 2,4-diacetylphloroglucinol resistance and in general stress tolerance. Microbiology 150:244-2450. - PubMed
-
- Baehler, E., M. Bottiglieri, M. Péchy-Tarr, M. Maurhofer, and C. Keel. 2005. Use of green fluorescent protein-based reporters to monitor balanced production of antifungal compounds in the biocontrol agent Pseudomonas fluorescens CHA0. J. Appl. Microbiol. 99:24-38. - PubMed
-
- Bao, Y., D. P. Lies, H. Fu, and G. P. Roberts. 1991. An improved Tn7-based system for the single-copy insertion of cloned genes into chromosomes of gram-negative bacteria. Gene 109:167-168. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

