A role for the Rab6A' GTPase in the inactivation of the Mad2-spindle checkpoint
- PMID: 16395330
- PMCID: PMC1383512
- DOI: 10.1038/sj.emboj.7600929
A role for the Rab6A' GTPase in the inactivation of the Mad2-spindle checkpoint
Abstract
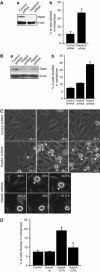
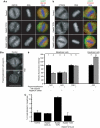
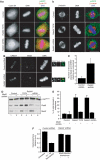
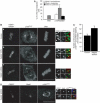
The two isoforms of the Rab6 GTPase, Rab6A and Rab6A', regulate a retrograde transport route connecting early endosomes and the endoplasmic reticulum via the Golgi complex in interphasic cells. Here we report that when Rab6A' function is altered cells are unable to progress normally through mitosis. Such cells are blocked in metaphase, despite displaying a normal Golgi fragmentation and with the Mad2-spindle checkpoint activated. Furthermore, the Rab6 effector p150(Glued), a subunit of the dynein/dynactin complex, remains associated with some kinetochores. A similar phenotype was observed when GAPCenA, a GTPase-activating protein of Rab6, was depleted from cells. Our results suggest that Rab6A' likely regulates the dynamics of the dynein/dynactin complex at the kinetochores and consequently the inactivation of the Mad2-spindle checkpoint. Rab6A', through its interaction with p150(Glued) and GAPCenA, may thus participate in a pathway involved in the metaphase/anaphase transition.
Figures
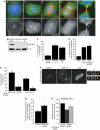
Similar articles
-
Rab6A and Rab6A' GTPases play non-overlapping roles in membrane trafficking.Traffic. 2006 Apr;7(4):394-407. doi: 10.1111/j.1600-0854.2006.00395.x. Traffic. 2006. PMID: 16536738
-
Regulation of microtubule-dependent recycling at the trans-Golgi network by Rab6A and Rab6A'.Mol Biol Cell. 2005 Jan;16(1):162-77. doi: 10.1091/mbc.e04-03-0260. Epub 2004 Oct 13. Mol Biol Cell. 2005. PMID: 15483056 Free PMC article.
-
A role for the Rab6B Bicaudal-D1 interaction in retrograde transport in neuronal cells.Exp Cell Res. 2007 Oct 1;313(16):3408-20. doi: 10.1016/j.yexcr.2007.05.032. Epub 2007 Jul 17. Exp Cell Res. 2007. PMID: 17707369
-
Transport of ricin from endosomes to the Golgi apparatus is regulated by Rab6A and Rab6A'.Traffic. 2006 Jun;7(6):663-72. doi: 10.1111/j.1600-0854.2006.00418.x. Traffic. 2006. PMID: 16683916
-
Ect2 and MgcRacGAP regulate the activation and function of Cdc42 in mitosis.J Cell Biol. 2005 Jan 17;168(2):221-32. doi: 10.1083/jcb.200408085. Epub 2005 Jan 10. J Cell Biol. 2005. PMID: 15642749 Free PMC article.
Cited by
-
Crystallization and preliminary X-ray crystallographic studies of Rab6A'(Q72L): a GTP-locked form.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012 Sep 1;68(Pt 9):1077-80. doi: 10.1107/S1744309112030874. Epub 2012 Aug 31. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012. PMID: 22949199 Free PMC article.
-
C11ORF24 is a novel type I membrane protein that cycles between the Golgi apparatus and the plasma membrane in Rab6-positive vesicles.PLoS One. 2013 Dec 2;8(12):e82223. doi: 10.1371/journal.pone.0082223. eCollection 2013. PLoS One. 2013. PMID: 24312644 Free PMC article.
-
Small GTPase Rab5 participates in chromosome congression and regulates localization of the centromere-associated protein CENP-F to kinetochores.Proc Natl Acad Sci U S A. 2011 Oct 18;108(42):17337-42. doi: 10.1073/pnas.1103516108. Epub 2011 Oct 10. Proc Natl Acad Sci U S A. 2011. PMID: 21987812 Free PMC article.
-
Electron tomography reveals Rab6 is essential to the trafficking of trans-Golgi clathrin and COPI-coated vesicles and the maintenance of Golgi cisternal number.Traffic. 2012 May;13(5):727-44. doi: 10.1111/j.1600-0854.2012.01343.x. Epub 2012 Mar 14. Traffic. 2012. PMID: 22335553 Free PMC article.
-
Cancer driver candidate genes AVL9, DENND5A and NUPL1 contribute to MDCK cystogenesis.Oncoscience. 2014 Dec 15;1(12):854-865. doi: 10.18632/oncoscience.107. eCollection 2014. Oncoscience. 2014. PMID: 25621300 Free PMC article.
References
-
- Antony C, Cibert C, Geraud G, Santa Maria A, Maro B, Mayau V, Goud B (1992) The small GTP-binding protein rab6p is distributed from medial Golgi to the trans-Golgi network as determined by a confocal microscopic approach. J Cell Sci 103: 785–796 - PubMed
-
- Aumais JP, Williams SN, Luo W, Nishino M, Caldwell KA, Caldwell GA, Lin SH, Yu-Lee LY (2003) Role for NudC, a dynein-associated nuclear movement protein, in mitosis and cytokinesis. J Cell Sci 116: 1991–2003 - PubMed
-
- Bailly E, McCaffrey M, Touchot N, Zahraoui A, Goud B, Bornens M (1991) Phosphorylation of two small GTP-binding proteins of the Rab family by p34cdc2. Nature 350: 715–718 - PubMed
-
- Bardin AJ, Amon A (2001) Men and sin: what's the difference? Nat Rev Mol Cell Biol 2: 815–826 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

