MSH2 is essential for the preservation of genome integrity and prevents homeologous recombination in the moss Physcomitrella patens
- PMID: 16397301
- PMCID: PMC1325206
- DOI: 10.1093/nar/gkj423
MSH2 is essential for the preservation of genome integrity and prevents homeologous recombination in the moss Physcomitrella patens
Abstract
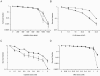
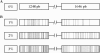
MSH2 is a central component of the mismatch repair pathway that targets mismatches arising during DNA replication, homologous recombination (HR) and in response to genotoxic stresses. Here, we describe the function of MSH2 in the moss Physcomitrella patens, as deciphered by the analysis of loss of function mutants. Ppmsh2 mutants display pleiotropic growth and developmental defects, which reflect genomic instability. Based on loss of function of the APT gene, we estimated this mutator phenotype to be at least 130 times higher in the mutants than in wild type. We also found that MSH2 is involved in some but not all the moss responses to genotoxic stresses we tested. Indeed, the Ppmsh2 mutants were more tolerant to cisplatin and show higher sensitivity to UV-B radiations. PpMSH2 gene involvement in HR was studied by assessing gene targeting (GT) efficiency with homologous and homeologous sequences. GT efficiency with homologous sequences was slightly decreased in the Ppmsh2 mutant compared with wild type. Strikingly GT efficiency with homeologous sequences decreased proportionally to sequence divergence in the wild type whereas it remained unaffected in the mutants. Those results demonstrate the role of PpMSH2 in the maintenance of genome integrity and in homologous and homeologous recombination.
Figures


Similar articles
-
Cloning of the PpMSH-2 cDNA of Physcomitrella patens, a moss in which gene targeting by homologous recombination occurs at high frequency.Biochimie. 2001 Nov-Dec;83(11-12):1003-8. doi: 10.1016/s0300-9084(01)01350-5. Biochimie. 2001. PMID: 11879728
-
RAD51 loss of function abolishes gene targeting and de-represses illegitimate integration in the moss Physcomitrella patens.DNA Repair (Amst). 2010 May 4;9(5):526-33. doi: 10.1016/j.dnarep.2010.02.001. Epub 2010 Mar 1. DNA Repair (Amst). 2010. PMID: 20189889
-
RECA plays a dual role in the maintenance of chloroplast genome stability in Physcomitrella patens.Plant J. 2015 Nov;84(3):516-26. doi: 10.1111/tpj.13017. Epub 2015 Oct 15. Plant J. 2015. PMID: 26340426
-
[The moss Physcomitrella patens, a new model system for functional genomics].Yi Chuan. 2004 Jul;26(4):560-6. Yi Chuan. 2004. PMID: 15640062 Review. Chinese.
-
The moss Physcomitrella patens.Annu Rev Genet. 2005;39:339-58. doi: 10.1146/annurev.genet.39.073003.110214. Annu Rev Genet. 2005. PMID: 16285864 Review.
Cited by
-
Repair of DNA double-strand breaks in plant meiosis: role of eukaryotic RecA recombinases and their modulators.Plant Reprod. 2023 Mar;36(1):17-41. doi: 10.1007/s00497-022-00443-6. Epub 2022 Jun 1. Plant Reprod. 2023. PMID: 35641832 Review.
-
Genotoxin induced mutagenesis in the model plant Physcomitrella patens.Biomed Res Int. 2013;2013:535049. doi: 10.1155/2013/535049. Epub 2013 Dec 9. Biomed Res Int. 2013. PMID: 24383055 Free PMC article.
-
MRE11 and RAD50, but not NBS1, are essential for gene targeting in the moss Physcomitrella patens.Nucleic Acids Res. 2012 Apr;40(8):3496-510. doi: 10.1093/nar/gkr1272. Epub 2011 Dec 30. Nucleic Acids Res. 2012. PMID: 22210882 Free PMC article.
-
CRISPR-Cas9-mediated efficient directed mutagenesis and RAD51-dependent and RAD51-independent gene targeting in the moss Physcomitrella patens.Plant Biotechnol J. 2017 Jan;15(1):122-131. doi: 10.1111/pbi.12596. Epub 2016 Jul 22. Plant Biotechnol J. 2017. PMID: 27368642 Free PMC article.
-
Genetic analysis of DEFECTIVE KERNEL1 loop function in three-dimensional body patterning in Physcomitrella patens.Plant Physiol. 2014 Oct;166(2):903-19. doi: 10.1104/pp.114.243758. Epub 2014 Sep 2. Plant Physiol. 2014. PMID: 25185121 Free PMC article.
References
-
- Modrich P., Lahue R. Mismatch repair in replication fidelity, genetic recombination, and cancer biology. Annu. Rev. Biochem. 1996;65:101–133. - PubMed
-
- Harfe B.D., Jinks-Robertson S. DNA mismatch repair and genetic instability. Annu. Rev. Genet. 2000;34:359–399. - PubMed
-
- Kolodner R. Biochemistry and genetics of eukaryotic mismatch repair. Genes Dev. 1996;10:1433–1442. - PubMed
-
- Schofield M.J., Hsieh P. DNA mismatch repair: molecular mechanisms and biological function. Annu. Rev. Microbiol. 2003;57:579–608. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

